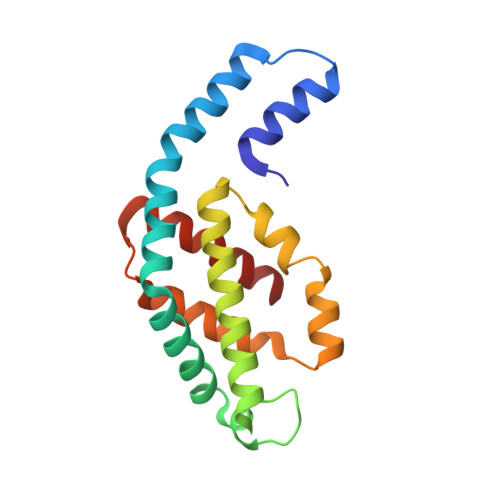
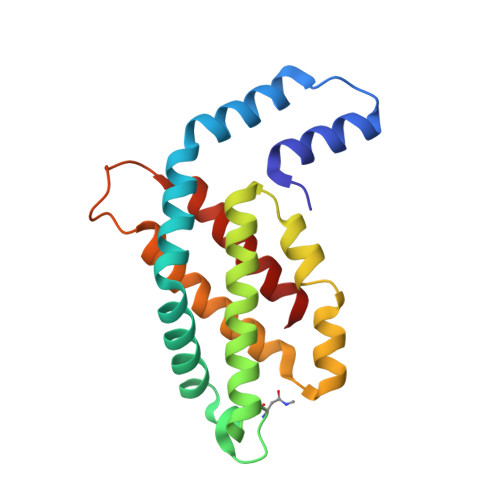
C-phycocyanin as a highly attractive model system in protein crystallography: unique crystallization properties and packing-diversity screening.
Sarrou, I., Feiler, C.G., Falke, S., Peard, N., Yefanov, O., Chapman, H.(2021) Acta Crystallogr D Struct Biol 77: 224-236
- PubMed: 33559611
- DOI: https://doi.org/10.1107/S2059798320016071
- Primary Citation of Related Structures:
6YPQ, 6YQ8, 6YQG, 6YYJ - PubMed Abstract:
The unique crystallization properties of the antenna protein C-phycocyanin (C-PC) from the thermophilic cyanobacterium Thermosynechococcus elongatus are reported and discussed. C-PC crystallizes in hundreds of significantly different conditions within a broad pH range and in the presence of a wide variety of precipitants and additives. Remarkably, the crystal dimensions vary from a few micrometres, as used in serial crystallography, to several hundred micrometres, with a very diverse crystal morphology. More than 100 unique single-crystal X-ray diffraction data sets were collected from randomly selected crystals and analysed. The addition of small-molecule additives revealed three new crystal packings of C-PC, which are discussed in detail. The high propensity of this protein to crystallize, combined with its natural blue colour and its fluorescence characteristics, make it an excellent candidate as a superior and highly adaptable model system in crystallography. C-PC can be used in technical and methods development approaches for X-ray and neutron diffraction techniques, and as a system for comprehending the fundamental principles of protein crystallography.
Organizational Affiliation:
Centre for Free-Electron Laser Science, DESY, Notkestrasse 85, 22607 Hamburg, Germany.