Activation and inhibition mechanisms of a plant helper NLR
Xiao, Y., Wu, X., Wang, Z., Wan, L.(2025) Nature
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Protein EDS1 | 623 | Arabidopsis thaliana | Mutation(s): 0 Gene Names: EDS1, EDS1-90, EDS1A, At3g48090, T17F15.40 | 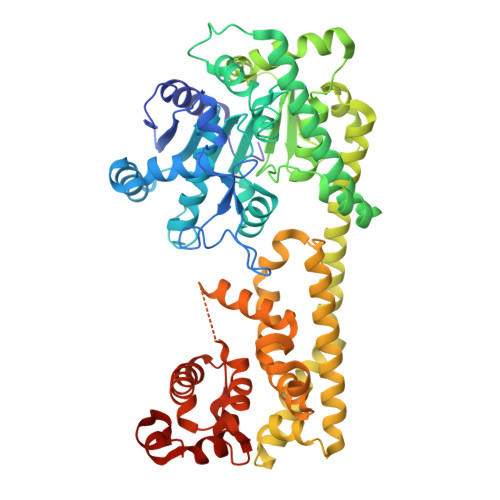 | |
UniProt | |||||
Find proteins for Q9SU72 (Arabidopsis thaliana) Explore Q9SU72 Go to UniProtKB: Q9SU72 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9SU72 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Senescence-associated carboxylesterase 101 | 537 | Arabidopsis thaliana | Mutation(s): 0 Gene Names: SAG101, At5g14930, F2G14.50 EC: 3.1.1.1 | 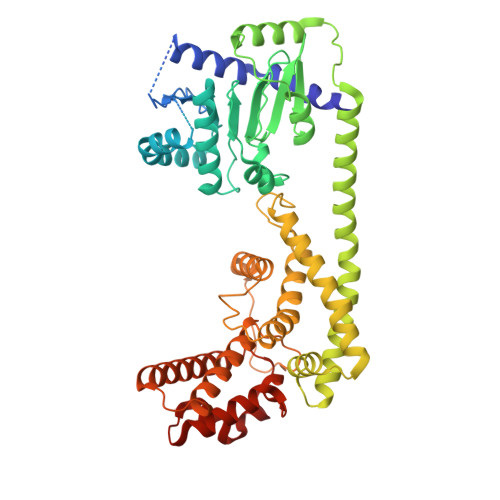 | |
UniProt | |||||
Find proteins for Q4F883 (Arabidopsis thaliana) Explore Q4F883 Go to UniProtKB: Q4F883 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q4F883 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Probable disease resistance protein At5g66890 | 415 | Arabidopsis thaliana | Mutation(s): 0 Gene Names: At5g66890, MUD21.15 | 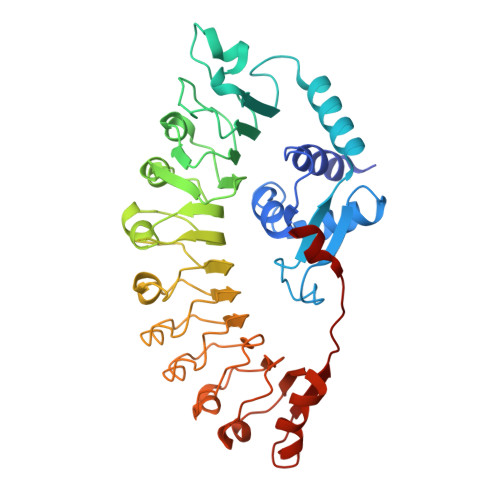 | |
UniProt | |||||
Find proteins for Q9FKZ2 (Arabidopsis thaliana) Explore Q9FKZ2 Go to UniProtKB: Q9FKZ2 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9FKZ2 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| A1L15 (Subject of Investigation/LOI) Query on A1L15 | D [auth A] | [[(2~{R},3~{R},4~{R},5~{R})-5-(6-aminopurin-9-yl)-4-[(2~{S},3~{R},4~{S},5~{R})-5-[[[[(2~{R},3~{S},4~{R},5~{R})-5-(6-aminopurin-9-yl)-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl]oxy-oxidanyl-phosphoryl]oxymethyl]-3,4-bis(oxidanyl)oxolan-2-yl]oxy-3-oxidanyl-oxolan-2-yl]methoxy-oxidanyl-phosphoryl] phosphono hydrogen phosphate C25 H37 N10 O26 P5 BLHBPWIRUPJXKW-VIUUMBSUSA-N |  | ||
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 32270304 |
| Chinese Academy of Sciences | China | XDB27040214 |