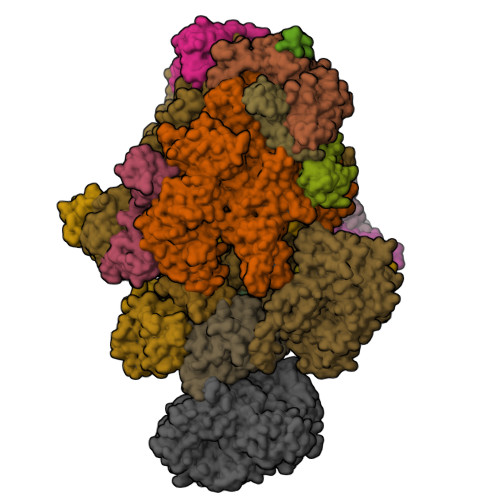
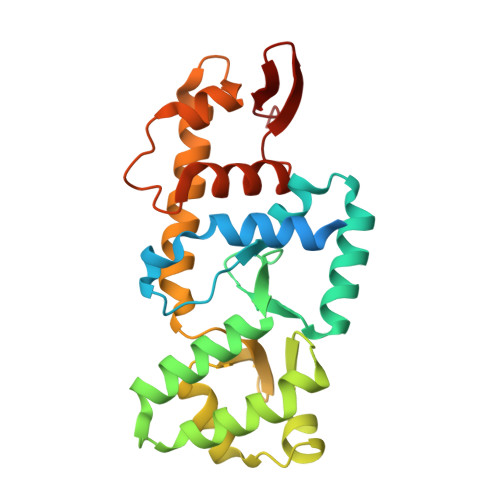
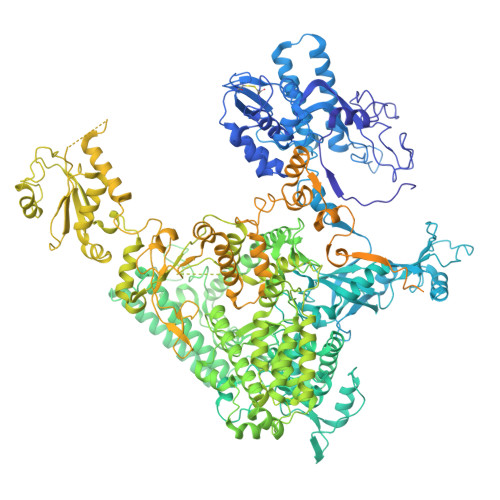
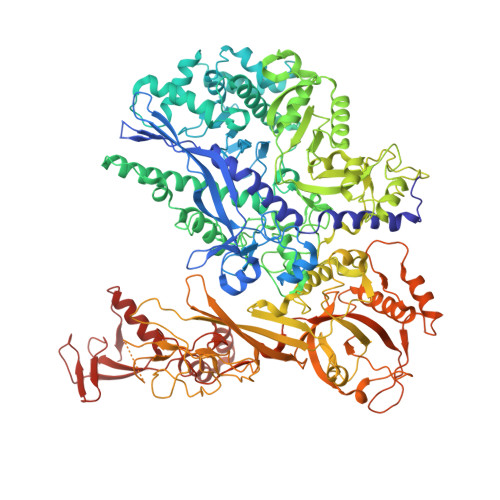
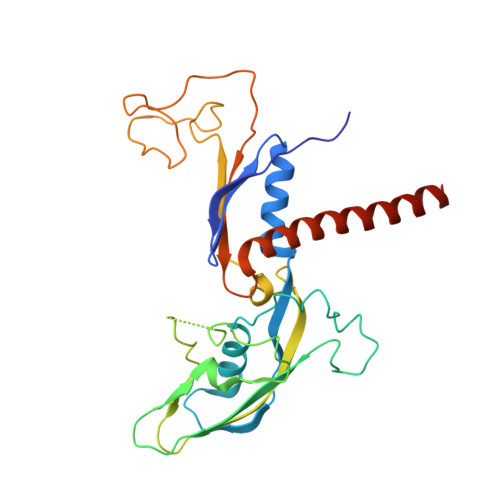
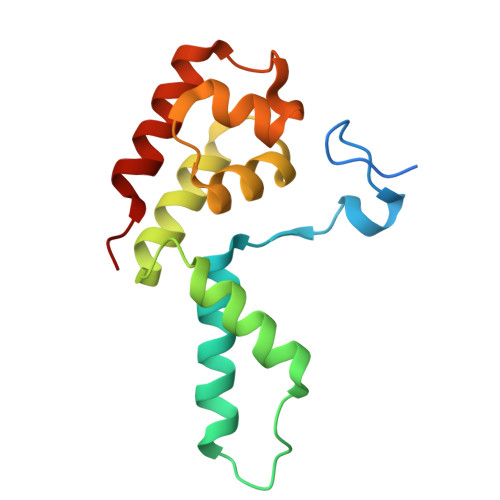
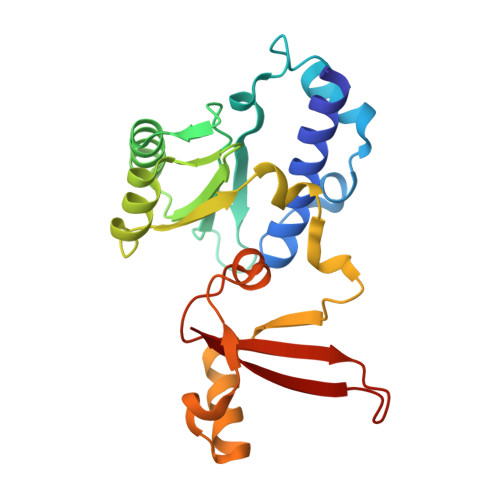
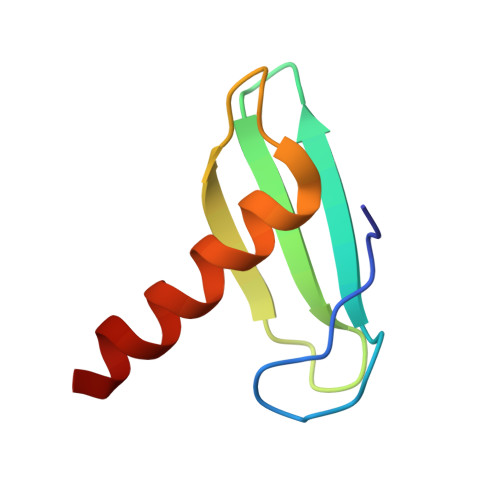
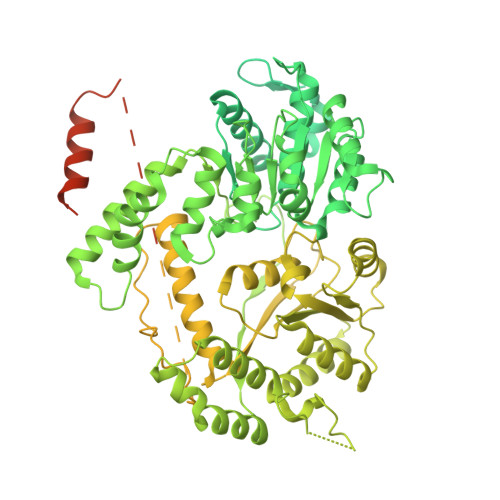
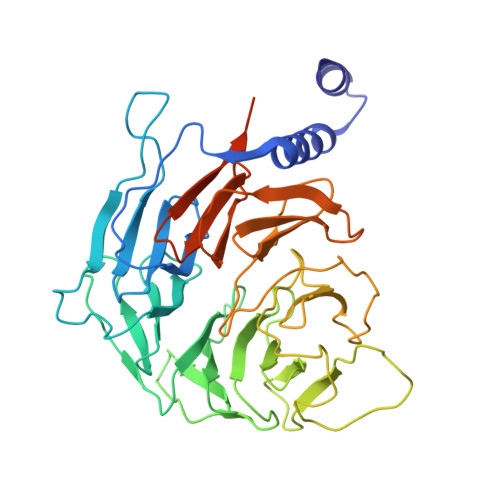
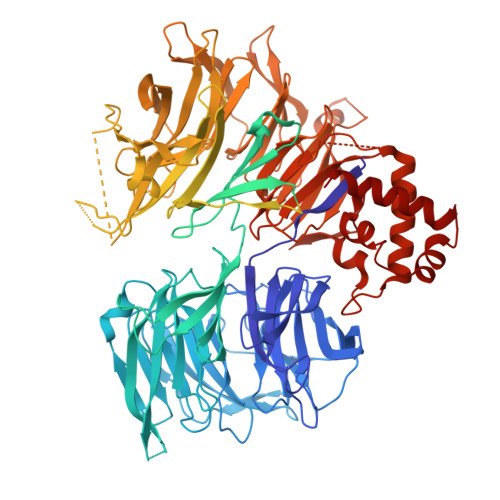
STK19 facilitates the clearance of lesion-stalled RNAPII during transcription-coupled DNA repair.
van den Heuvel, D., Rodriguez-Martinez, M., van der Meer, P.J., Nieto Moreno, N., Park, J., Kim, H.S., van Schie, J.J.M., Wondergem, A.P., D'Souza, A., Yakoub, G., Herlihy, A.E., Kashyap, K., Boissiere, T., Walker, J., Mitter, R., Apelt, K., de Lint, K., Kirdok, I., Ljungman, M., Wolthuis, R.M.F., Cramer, P., Scharer, O.D., Kokic, G., Svejstrup, J.Q., Luijsterburg, M.S.(2024) Cell 187: 7107
- PubMed: 39547229
- DOI: https://doi.org/10.1016/j.cell.2024.10.018
- Primary Citation of Related Structures:
9ER2 - PubMed Abstract:
Transcription-coupled DNA repair (TCR) removes bulky DNA lesions impeding RNA polymerase II (RNAPII) transcription. Recent studies have outlined the stepwise assembly of TCR factors CSB, CSA, UVSSA, and transcription factor IIH (TFIIH) around lesion-stalled RNAPII. However, the mechanism and factors required for the transition to downstream repair steps, including RNAPII removal to provide repair proteins access to the DNA lesion, remain unclear. Here, we identify STK19 as a TCR factor facilitating this transition. Loss of STK19 does not impact initial TCR complex assembly or RNAPII ubiquitylation but delays lesion-stalled RNAPII clearance, thereby interfering with the downstream repair reaction. Cryoelectron microscopy (cryo-EM) and mutational analysis reveal that STK19 associates with the TCR complex, positioning itself between RNAPII, UVSSA, and CSA. The structural insights and molecular modeling suggest that STK19 positions the ATPase subunits of TFIIH onto DNA in front of RNAPII. Together, these findings provide new insights into the factors and mechanisms required for TCR.
Organizational Affiliation:
Department of Human Genetics, Leiden University Medical Center, Leiden, the Netherlands.