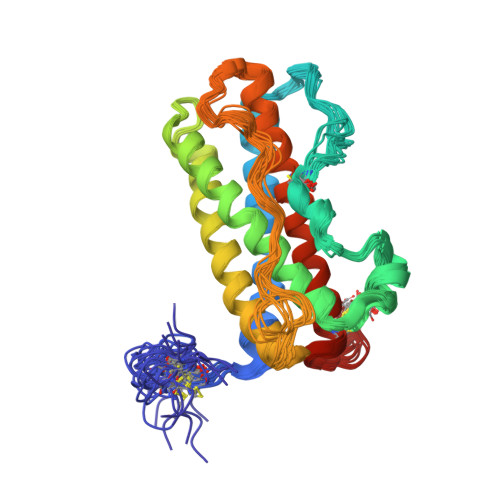
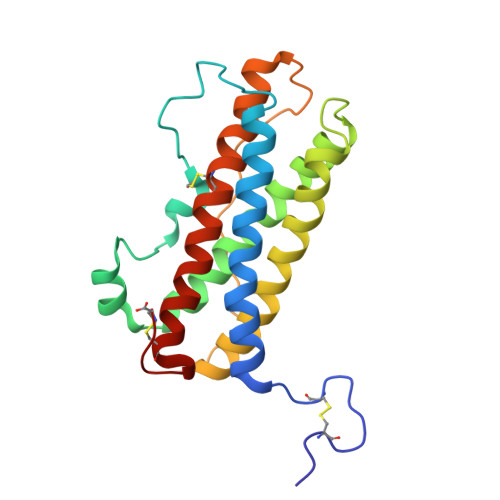
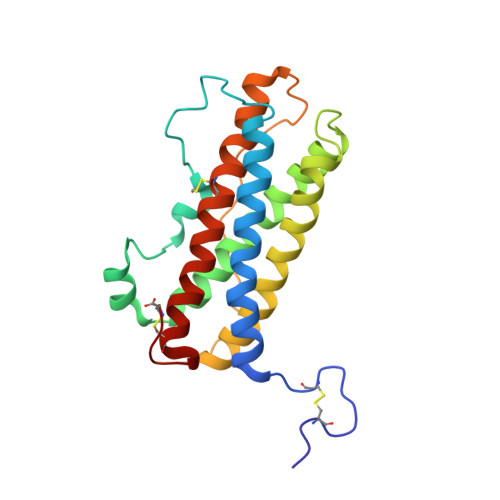
Solution structure of human prolactin
Teilum, K., Hoch, J.C., Goffin, V., Kinet, S., Martial, J.A., Kragelund, B.B.(2005) J Mol Biology 351: 810-823
- PubMed: 16045928
- DOI: https://doi.org/10.1016/j.jmb.2005.06.042
- Primary Citation of Related Structures:
1RW5 - PubMed Abstract:
We report the solution structure of human prolactin determined by NMR spectroscopy. Our result is a significant improvement over a previous structure in terms of number and distribution of distance restraints, regularity of secondary structure, and potential energy. More significantly, the structure is sufficiently different that it leads to different conclusions regarding the mechanism of receptor activation and initiation of signal transduction. Here, we compare the structure of unbound prolactin to structures of both the homologue ovine placental lactogen and growth hormone. The structures of unbound and receptor bound prolactin/placental lactogen are similar and no noteworthy structural changes occur upon receptor binding. The observation of enhanced binding at the second receptor site when the first site is occupied has been widely interpreted to indicate conformational change induced by binding the first receptor. However, our results indicate that this enhanced binding at the second site could be due to receptor-receptor interactions or some other free energy sources rather than conformational change in the hormone. Titration of human prolactin with the extracellular domain of the human prolactin receptor was followed by NMR, gel filtration and electrophoresis. Both binary and ternary hormone-receptor complexes are clearly detectable by gel filtration and electrophoresis. The binary complex is not observable by NMR, possibly due to a dynamic equilibrium in intermediate exchange within the complex. The ternary complex of one hormone molecule bound to two receptor molecules is on the contrary readily detectable by NMR. This is in stark contrast to the widely held view that the ternary prolactin-receptor complex is only transiently formed. Thus, our results lead to improved understanding of the prolactin-prolactin receptor interaction.
Organizational Affiliation:
Department of Protein Chemistry, Institute of Molecular Biology and Physiology, University of Copenhagen, Øster Farimagsgade 2A, DK-1353 Copenhagen K, Denmark.