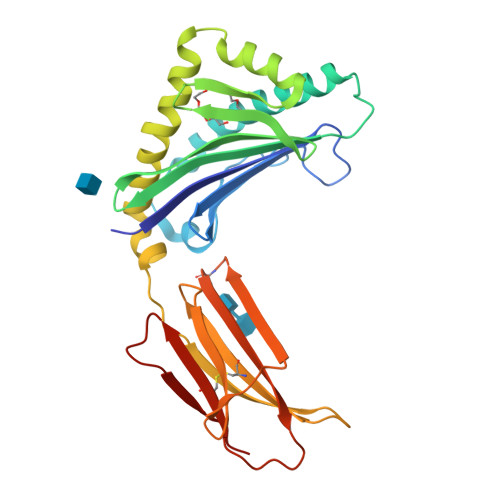
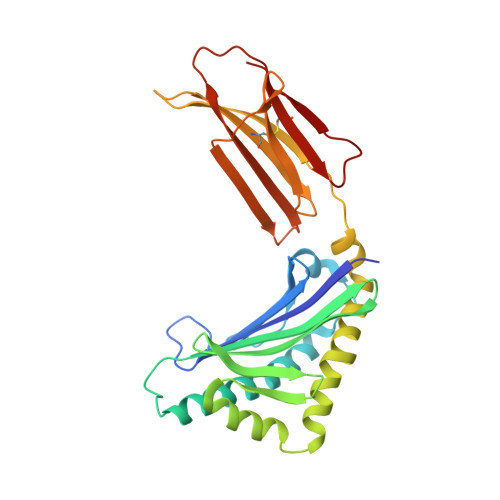
Crystallographic studies of ligand binding by Zn-alpha2-glycoprotein.
Delker, S.L., West Jr., A.P., McDermott, L., Kennedy, M.W., Bjorkman, P.J.(2004) J Struct Biol 148: 205-213
- PubMed: 15477100
- DOI: https://doi.org/10.1016/j.jsb.2004.04.009
- Primary Citation of Related Structures:
1T7V, 1T7W, 1T7X, 1T7Y, 1T7Z, 1T80 - PubMed Abstract:
Zn-alpha2-glycoprotein (ZAG) is a 41 kDa soluble protein that is present in most bodily fluids. The previously reported 2.8 A crystal structure of ZAG isolated from human serum demonstrated the structural similarity between ZAG and class I major histocompatibility complex (MHC) molecules and revealed a non-peptidic ligand in the ZAG counterpart of the MHC peptide-binding groove. Here we present crystallographic studies to explore further the nature of the non-peptidic ligand in the ZAG groove. Comparison of the structures of several forms of recombinant ZAG, including a 1.95 A structure derived from ZAG expressed in insect cells, suggests that the non-peptidic ligand in the current structures and in the structure of serum ZAG is a polyethylene glycol (PEG), which is present in the crystallization conditions used. Further support for PEG binding in the ZAG groove is provided by the finding that PEG displaces a fluorophore-tagged fatty acid from the ZAG binding site. From these results we hypothesize that our purified forms of ZAG do not contain a bound endogenous ligand, but that the ZAG groove is capable of binding hydrophobic molecules, which may relate to its function.
Organizational Affiliation:
Division of Biology, 114-96 Pasadena, CA 91125, USA.