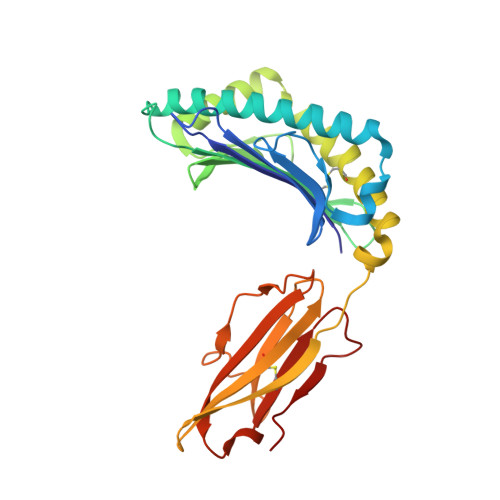
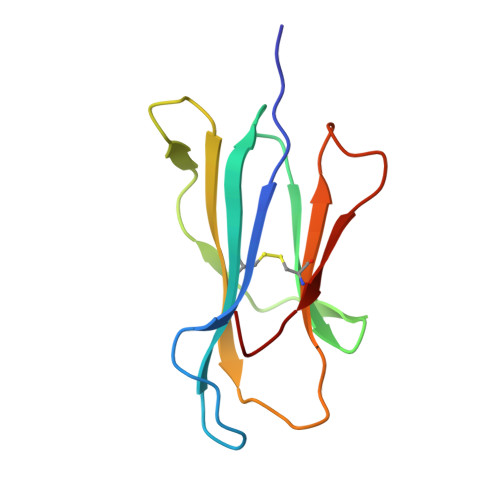
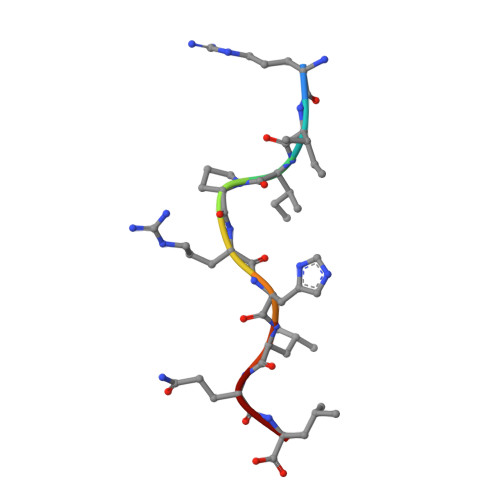
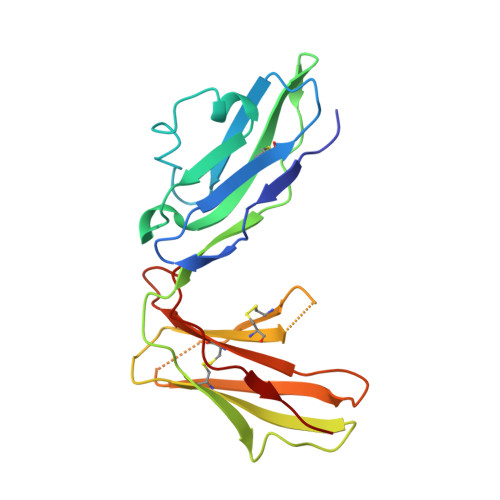
Structural basis for recognition of the nonclassical MHC molecule HLA-G by the leukocyte Ig-like receptor B2 (LILRB2/LIR2/ILT4/CD85d)
Shiroishi, M., Kuroki, K., Rasubala, L., Tsumoto, K., Kumagai, I., Kurimoto, E., Kato, K., Kohda, D., Maenaka, K.(2006) Proc Natl Acad Sci U S A 103: 16412-16417
- PubMed: 17056715
- DOI: https://doi.org/10.1073/pnas.0605228103
- Primary Citation of Related Structures:
2DYP - PubMed Abstract:
HLA-G is a nonclassical MHC class I (MHCI) molecule that can suppress a wide range of immune responses in the maternal-fetal interface. The human inhibitory immune receptors leukocyte Ig-like receptor (LILR) B1 [also called LIR1, Ig-like transcript 2 (ILT2), or CD85j] and LILRB2 (LIR2/ILT4/CD85d) preferentially recognize HLA-G. HLA-G inherently exhibits various forms, including beta(2)-microglobulin (beta(2)m)-free and disulfide-linked dimer forms. Notably, LILRB1 cannot recognize the beta(2)m-free form of HLA-G or HLA-B27, but LILRB2 can recognize the beta(2)m-free form of HLA-B27. To date, the structural basis for HLA-G/LILR recognition remains to be examined. Here, we report the 2.5-A resolution crystal structure of the LILRB2/HLA-G complex. LILRB2 exhibits an overlapping but distinct MHCI recognition mode compared with LILRB1 and dominantly recognizes the hydrophobic site of the HLA-G alpha3 domain. NMR binding studies also confirmed these LILR recognition differences on both conformed (heavy chain/peptide/beta(2)m) and free forms of beta(2)m. Binding studies using beta(2)m-free MHCIs revealed differential beta(2)m-dependent LILR-binding specificities. These results suggest that subtle structural differences between LILRB family members cause the distinct binding specificities to various forms of HLA-G and other MHCIs, which may in turn regulate immune suppression.
Organizational Affiliation:
Division of Structural Biology, Medical Institute of Bioregulation, Kyushu University, 3-1-1 Maidashi, Higashi-ku, Fukuoka 812-8582, Japan.