An Auto-Inhibitory Tyrosine Motif in the Cell-Cycle Regulated Nek7 Kinase is Released Through Binding of Nek9
Richards, M.W., O'Regan, L., Mas-Droux, C., Blot, J.M.Y., Cheung, J., Hoelder, S., Fry, A.M., Bayliss, R.(2009) Mol Cell 36: 560
- PubMed: 19941817
- DOI: https://doi.org/10.1016/j.molcel.2009.09.038
- Primary Citation of Related Structures:
2WQM, 2WQN, 2WQO - PubMed Abstract:
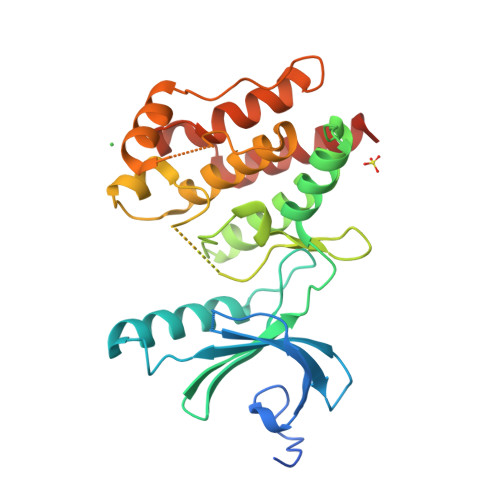
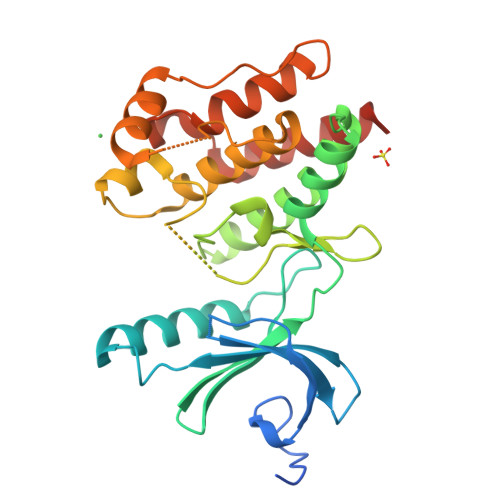
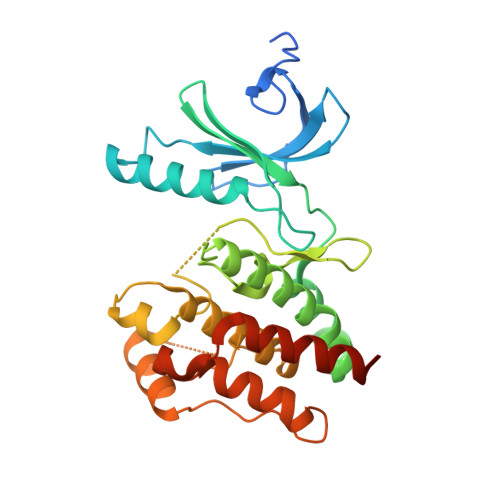
Mitosis is controlled by multiple protein kinases, many of which are abnormally expressed in human cancers. Nek2, Nek6, Nek7, and Nek9 are NIMA-related kinases essential for proper mitotic progression. We determined the atomic structure of Nek7 and discovered an autoinhibited conformation that suggests a regulatory mechanism not previously described in kinases. Additionally, Nek2 adopts the same conformation when bound to a drug-like molecule. In both structures, a tyrosine side chain points into the active site, interacts with the activation loop, and blocks the alphaC helix. Tyrosine mutants of Nek7 and the related kinase Nek6 are constitutively active. The activity of Nek6 and Nek7, but not the tyrosine mutant, is increased by interaction with the Nek9 noncatalytic C-terminal domain, suggesting a mechanism in which the tyrosine is released from its autoinhibitory position. The autoinhibitory conformation is common to three Neks and provides a potential target for selective kinase inhibitors.
Organizational Affiliation:
Institute of Cancer Research, London, UK.