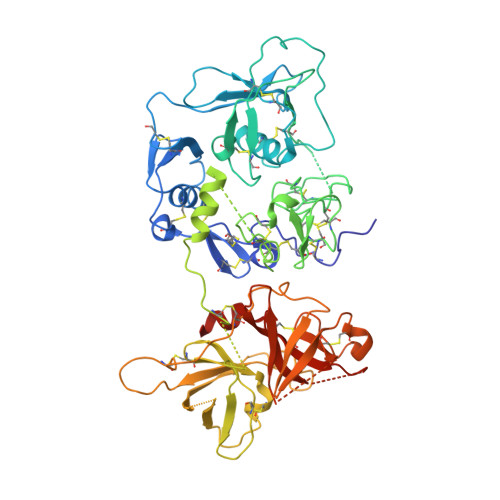
Structural Basis for Complement Factor I Control and its Disease-Associated Sequence Polymorphisms.
Roversi, P., Johnson, S., Caesar, J.J., Mclean, F., Leath, K.J., Tsiftsoglou, S.A., Morgan, B.P., Harris, C.L., Sim, R.B., Lea, S.M.(2011) Proc Natl Acad Sci U S A 108: 12839
- PubMed: 21768352
- DOI: https://doi.org/10.1073/pnas.1102167108
- Primary Citation of Related Structures:
2XRC - PubMed Abstract:
The complement system is a key component of innate and adaptive immune responses. Complement regulation is critical for prevention and control of disease. We have determined the crystal structure of the complement regulatory enzyme human factor I (fI). FI is in a proteolytically inactive form, demonstrating that it circulates in a zymogen-like state despite being fully processed to the mature sequence. Mapping of functional data from mutants of fI onto the structure suggests that this inactive form is maintained by the noncatalytic heavy-chain allosterically modulating activity of the light chain. Once the ternary complex of fI, a cofactor and a substrate is formed, the allosteric inhibition is released, and fI is oriented for cleavage. In addition to explaining how circulating fI is limited to cleaving only C3b/C4b, our model explains the molecular basis of disease-associated polymorphisms in fI and its cofactors.
Organizational Affiliation:
Sir William Dunn School of Pathology, University of Oxford, Oxford OX1 3RE, United Kingdom.