Class I Hdacs Share a Common Mechanism of Regulation by Inositol Phosphates.
Millard, C.J., Watson, P.J., Celardo, I., Gordiyenko, Y., Cowley, S.M., Robinson, C.V., Fairall, L., Schwabe, J.W.R.(2013) Mol Cell 51: 57
- PubMed: 23791785
- DOI: https://doi.org/10.1016/j.molcel.2013.05.020
- Primary Citation of Related Structures:
4BKX - PubMed Abstract:
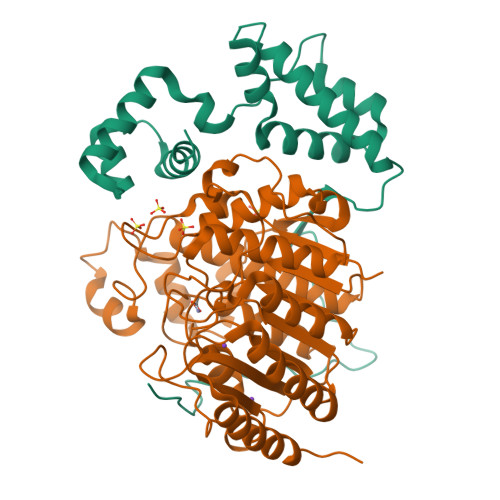
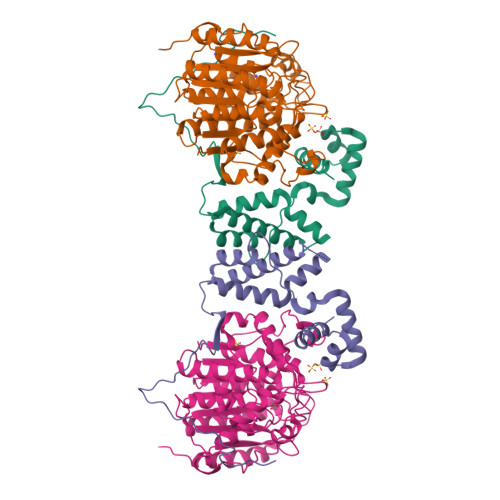
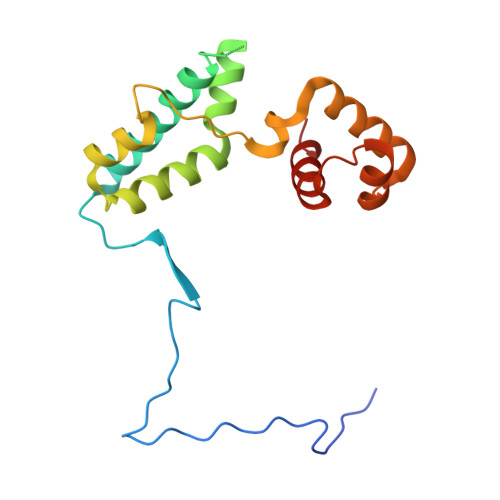
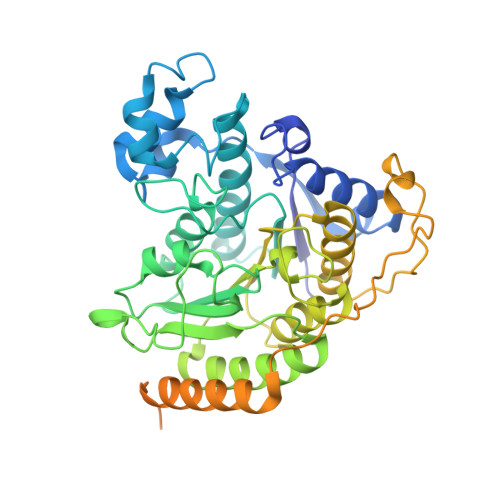
Class I histone deacetylases (HDAC1, HDAC2, and HDAC3) are recruited by cognate corepressor proteins into specific transcriptional repression complexes that target HDAC activity to chromatin resulting in chromatin condensation and transcriptional silencing. We previously reported the structure of HDAC3 in complex with the SMRT corepressor. This structure revealed the presence of inositol-tetraphosphate [Ins(1,4,5,6)P4] at the interface of the two proteins. It was previously unclear whether the role of Ins(1,4,5,6)P4 is to act as a structural cofactor or a regulator of HDAC3 activity. Here we report the structure of HDAC1 in complex with MTA1 from the NuRD complex. The ELM2-SANT domains from MTA1 wrap completely around HDAC1 occupying both sides of the active site such that the adjacent BAH domain is ideally positioned to recruit nucleosomes to the active site of the enzyme. Functional assays of both the HDAC1 and HDAC3 complexes reveal that Ins(1,4,5,6)P4 is a bona fide conserved regulator of class I HDAC complexes.
Organizational Affiliation:
Henry Wellcome Laboratories of Structural Biology, Department of Biochemistry, University of Leicester, UK.