Biological and Structural Evaluation of 10R- and 10S-Methylthio-DDACTHF Reveals a New Role for Sulfur in Inhibition of Glycinamide Ribonucleotide Transformylase.
Connelly, S., Demartino, J.K., Boger, D.L., Wilson, I.A.(2013) Biochemistry 52: 5133-5144
- PubMed: 23869564
- DOI: https://doi.org/10.1021/bi4005182
- Primary Citation of Related Structures:
4EW1, 4EW2, 4EW3 - PubMed Abstract:
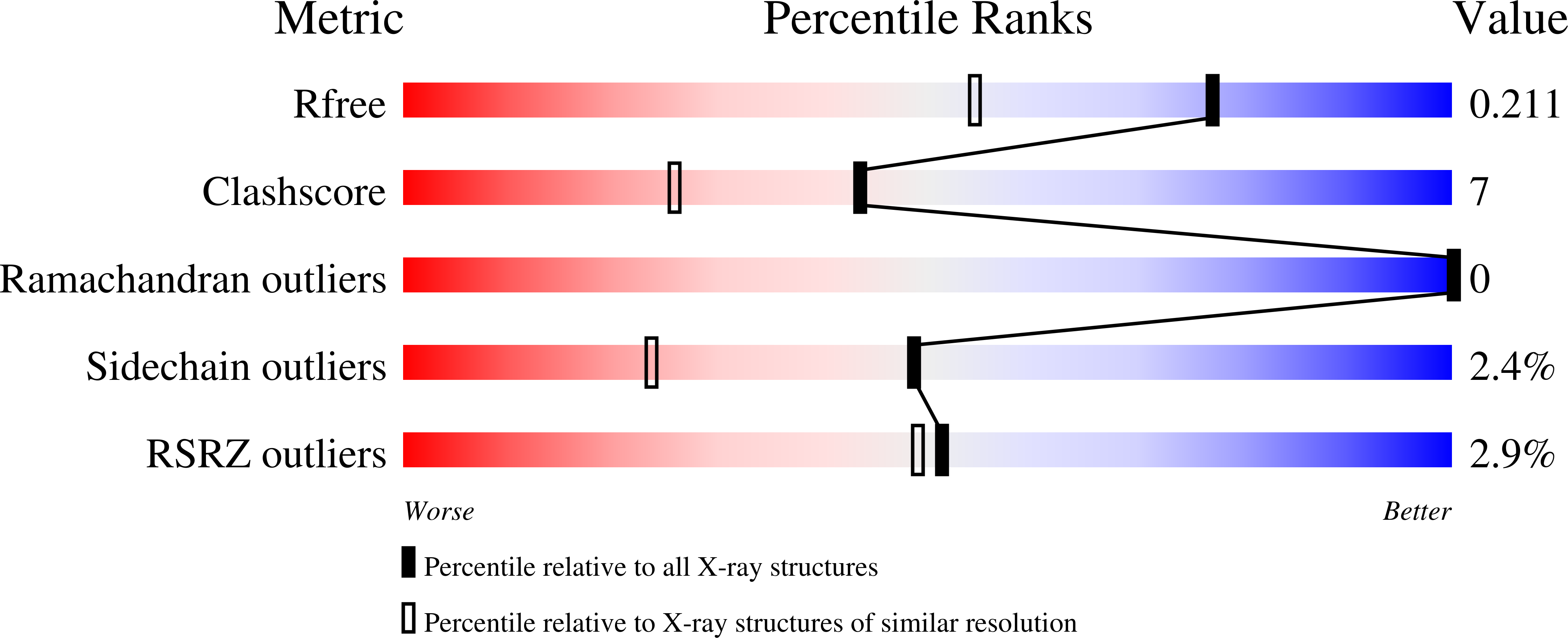
Glycinamide ribonucleotide transformylase (GAR Tfase) is a folate-dependent enzyme in the de novo purine biosynthesis pathway, which has long been considered a potential target for development of anti-neoplastic therapeutics. Here we report the biological and X-ray crystallographic evaluations of both independent C10 diastereomers, 10S- and 10R-methylthio-DDACTHF, bound to human GAR Tfase, including the highest-resolution apo GAR Tfase structure to date (1.52 Å). Both diastereomers are potent inhibitors (Ki = 210 nM for 10R, and Ki = 180 nM for 10S) of GAR Tfase and exhibit effective inhibition of human leukemia cell growth (IC₅₀ = 80 and 50 nM, respectively). Their inhibitory activity was surprisingly high, and these lipophilic C10-substituted analogues show distinct advantages over their hydrophilic counterparts, most strikingly in retaining potency in mutant human leukemia cell lines that lack reduced folate carrier protein activity (IC₅₀ = 70 and 60 nM, respectively). Structural characterization reveals a new binding mode for these diastereoisomers, in which the lipophilic thiomethyl groups penetrate deeper into a hydrophobic pocket within the folate-binding site. In silico docking simulations of three other sulfur-containing folate analogues also indicate that this hydrophobic cleft represents a favorable region for binding lipophilic substituents. Overall, these results suggest sulfur and its substitutions play an important role in not only the binding of anti-folates to GAR Tfase but also the selectivity and cellular activity (growth inhibition), thereby presenting new possibilities for the future design of potent and selective anti-folate drugs that target GAR Tfase.
Organizational Affiliation:
Department of Integrative Structural and Computational Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, California 92037, United States.