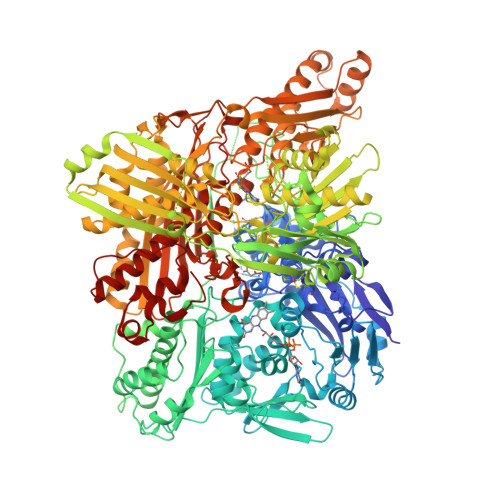
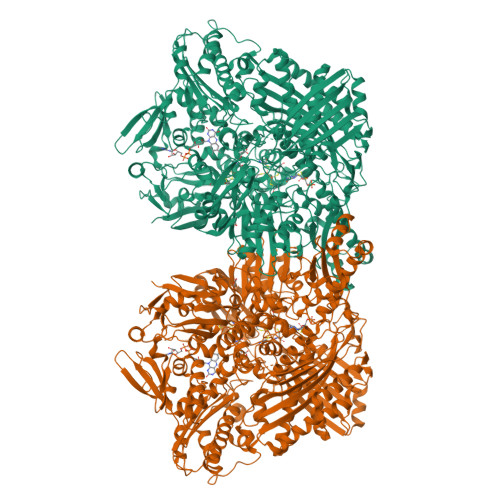
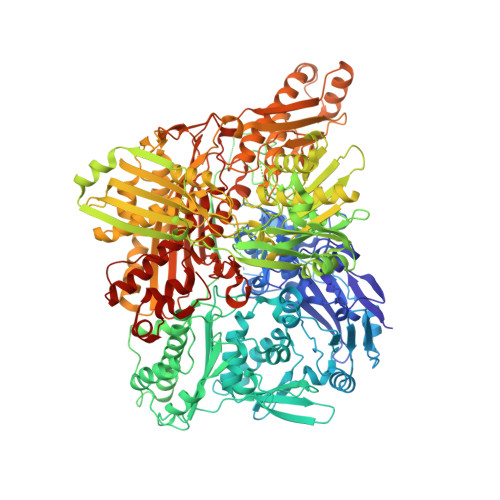
Structural Insights Into Xenobiotic and Inhibitor Binding to Human Aldehyde Oxidase
Coelho, C., Foti, A., Hartmann, T., Santos-Silva, T., Leimkuhler, S., Romao, M.J.(2015) Nat Chem Biol 11: 779
- PubMed: 26322824
- DOI: https://doi.org/10.1038/nchembio.1895
- Primary Citation of Related Structures:
4UHW, 4UHX - PubMed Abstract:
Aldehyde oxidase (AOX) is a xanthine oxidase (XO)-related enzyme with emerging importance due to its role in the metabolism of drugs and xenobiotics. We report the first crystal structures of human AOX1, substrate free (2.6-Å resolution) and in complex with the substrate phthalazine and the inhibitor thioridazine (2.7-Å resolution). Analysis of the protein active site combined with steady-state kinetic studies highlight the unique features, including binding and substrate orientation at the active site, that characterize human AOX1 as an important drug-metabolizing enzyme. Structural analysis of the complex with the noncompetitive inhibitor thioridazine revealed a new, unexpected and fully occupied inhibitor-binding site that is structurally conserved among mammalian AOXs and XO. The new structural insights into the catalytic and inhibition mechanisms of human AOX that we now report will be of great value for the rational analysis of clinical drug interactions involving inhibition of AOX1 and for the prediction and design of AOX-stable putative drugs.
Organizational Affiliation:
Research Unit on Applied Molecular Biosciences-Rede de Química e Tecnologia (UCIBIO-REQUIMTE), Departamento de Química, Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, Caparica, Portugal.