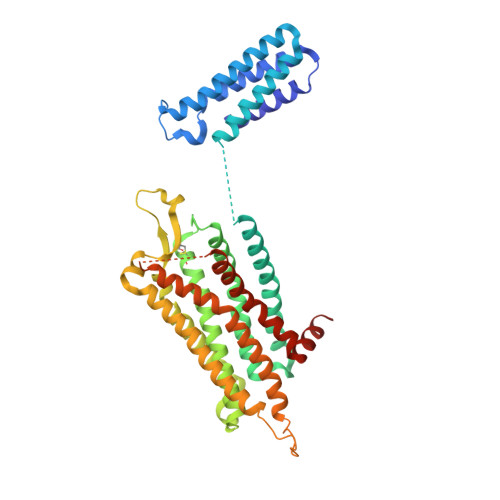
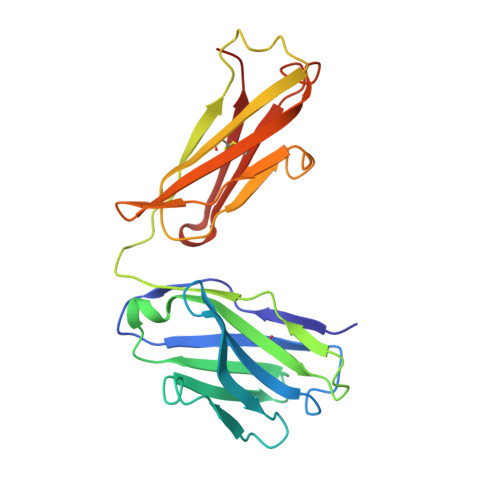
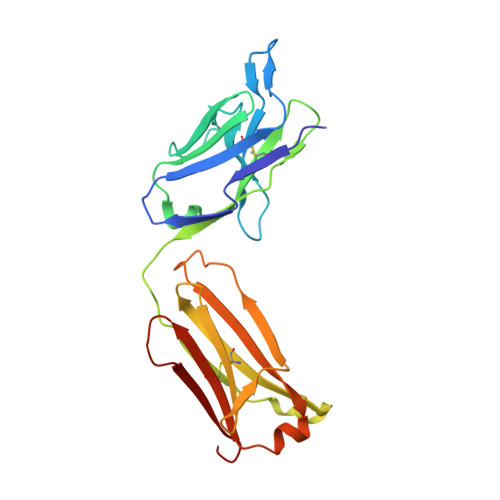
Structure of an antagonist-bound ghrelin receptor reveals possible ghrelin recognition mode.
Shiimura, Y., Horita, S., Hamamoto, A., Asada, H., Hirata, K., Tanaka, M., Mori, K., Uemura, T., Kobayashi, T., Iwata, S., Kojima, M.(2020) Nat Commun 11: 4160-4160
- PubMed: 32814772
- DOI: https://doi.org/10.1038/s41467-020-17554-1
- Primary Citation of Related Structures:
6KO5, 6KS2 - PubMed Abstract:
Ghrelin is a gastric peptide hormone with important physiological functions. The unique feature of ghrelin is its Serine 3 acyl-modification, which is essential for ghrelin's activity. However, it remains to be elucidated why the acyl-modification of ghrelin is necessary for activity. To address these questions, we solved the crystal structure of the ghrelin receptor bound to antagonist. The ligand-binding pocket of the ghrelin receptor is bifurcated by a salt bridge between E124 and R283. A striking feature of the ligand-binding pocket of the ghrelin receptor is a wide gap (crevasse) between the TM6 and TM7 bundles that is rich in hydrophobic amino acids, including a cluster of phenylalanine residues. Mutagenesis analyses suggest that the interaction between the gap structure and the acyl acid moiety of ghrelin may participate in transforming the ghrelin receptor into an active conformation.
Organizational Affiliation:
Division of Molecular Genetics, Institute of Life Science, Kurume University, Fukuoka, Japan.