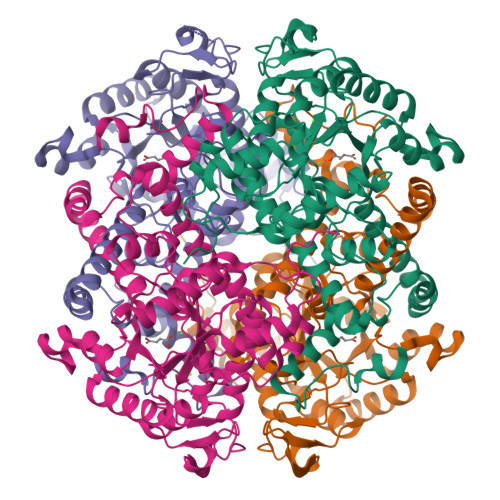
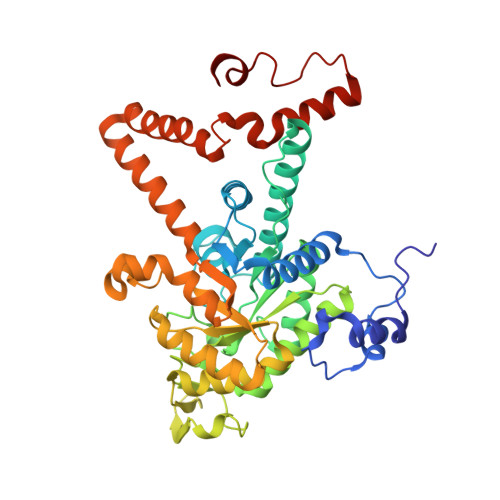
Itaconate is a covalent inhibitor of the Mycobacterium tuberculosis isocitrate lyase.
Kwai, B.X.C., Collins, A.J., Middleditch, M.J., Sperry, J., Bashiri, G., Leung, I.K.H.(2021) RSC Med Chem 12: 57-61
- PubMed: 34046597
- DOI: https://doi.org/10.1039/d0md00301h
- Primary Citation of Related Structures:
6XPP - PubMed Abstract:
Itaconate is a mammalian antimicrobial metabolite that inhibits the isocitrate lyases (ICLs) of Mycobacterium tuberculosis . Herein, we report that ICLs form a covalent adduct with itaconate through their catalytic cysteine residue. These results reveal atomic details of itaconate inhibition and provide insights into the catalytic mechanism of ICLs.
Organizational Affiliation:
School of Chemical Sciences, The University of Auckland Private Bag 92019, Victoria Street West Auckland 1142 New Zealand j.sperry@auckland.ac.nz i.leung@auckland.ac.nz.