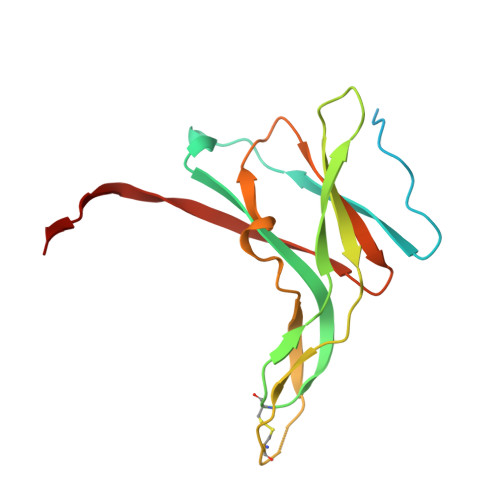
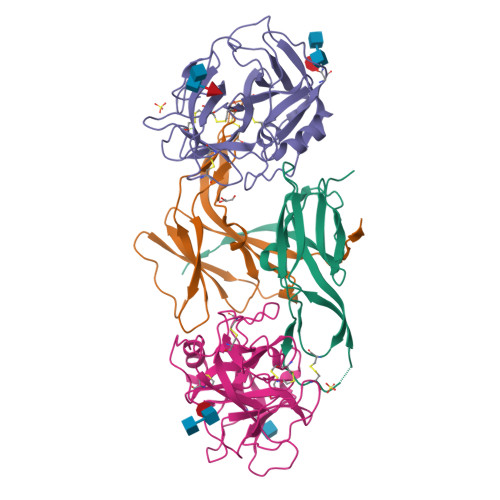
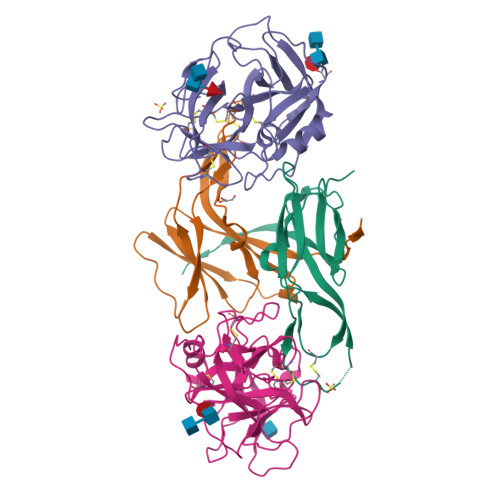
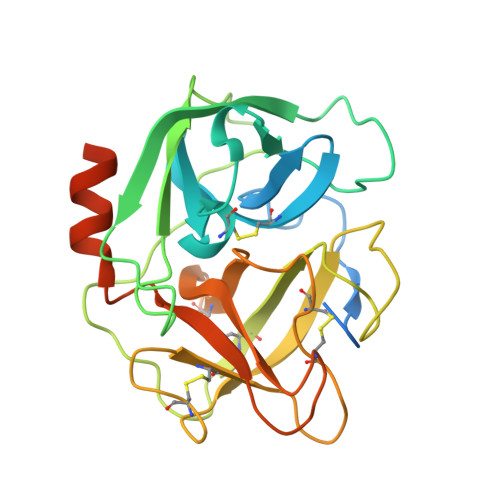
Structural Basis for the Inhibition Mechanism of Ecotin against Neutrophil Elastase by Targeting the Active Site and Secondary Binding Site.
Jobichen, C., Prabhakar, M.T., Loh, S.N., Sivaraman, J.(2020) Biochemistry 59: 2788-2795
- PubMed: 32657577
- DOI: https://doi.org/10.1021/acs.biochem.0c00493
- Primary Citation of Related Structures:
7CBK - PubMed Abstract:
Human neutrophil elastase (hNE) is a serine protease that plays a major role in defending the bacterial infection. However, elevated expression of hNE is reported in lung and breast cancer, among others. Moreover, hNE is a target for the treatment of cardiopulmonary diseases. Ecotin (ET) is a serine protease inhibitor present in many Gram-negative bacteria, and it plays a physiological role in inhibiting host proteases, including hNE. Despite this known interaction, the structure of the hNE-ET complex has not been reported, and the mechanism of ecotin inhibition is not available. We determined the structure of the hNE-ET complex by molecular replacement method. The structure of the hNE-ET complex revealed the presence of six interface regions comprising 50s, 60s, and 80s loops, between the ET dimer and two independent hNE monomers, which explains the high affinity of ecotin for hNE (12 pM). Notably, we observed a secondary binding site of hNE located 24 Å from the primary binding site. Comparison of the closely related trypsin-ecotin complex with our hNE-ET complex shows movement of the backbone atoms of the 80s and 50s loops by 4.6 Å, suggesting the flexibility of these loops in inhibiting a range of proteases. Through a detailed structural analysis, we demonstrate the flexibility of the hNE subsites to dock various side chains concomitant with inhibition, indicating the broad specificity of hNE against various inhibitors. These findings will aid in the design of chimeric inhibitors that target both sites of hNE and in the development of therapeutics for controlling hNE-mediated pathogenesis.
Organizational Affiliation:
Department of Biological Sciences, National University of Singapore, 14 Science Drive 4, Singapore 117543.