Full Text |
QUERY: Citation Author = "Yildirim, I." | MyPDB Login | Search API |
| Search Summary | This query matches 20 Structures. |
Structure Determination MethodologyScientific Name of Source OrganismTaxonomyExperimental MethodPolymer Entity TypeRefinement Resolution (Å)Release DateEnzyme Classification NameSymmetry Type | 1 to 20 of 20 Structures Page 1 of 1 Sort by
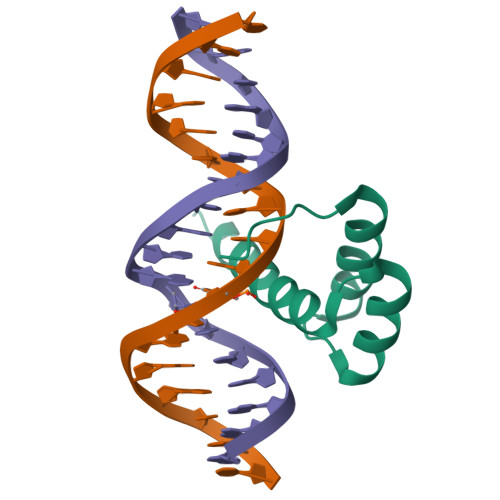
Structure of HOXB13 complex with methylated DNAMorgunova, E., Yin, Y., Jolma, A., Popov, A., Taipale, J. (2017) Science 356:
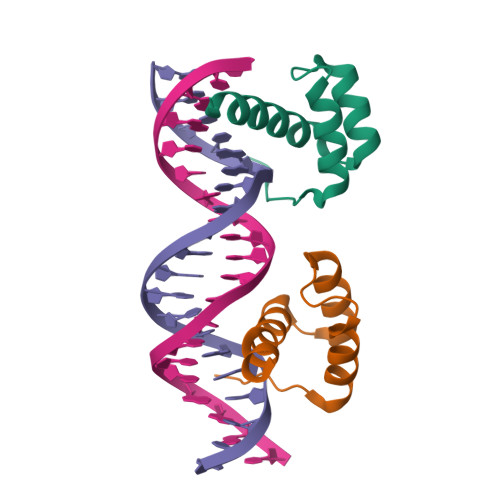
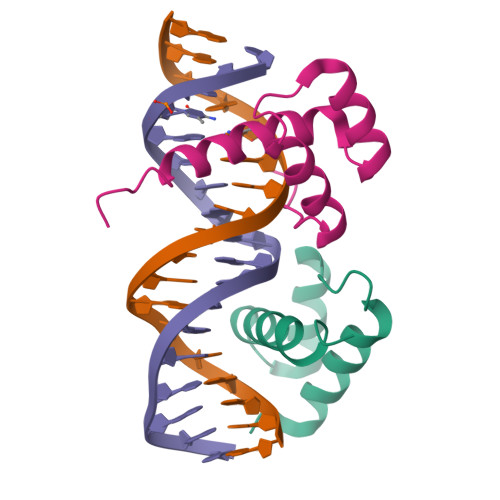
HOXB13-MEIS1 heterodimer bound to methylated DNAMorgunova, E., Yin, Y., Jolma, A., Popov, A., Taipale, J. (2017) Science 356:
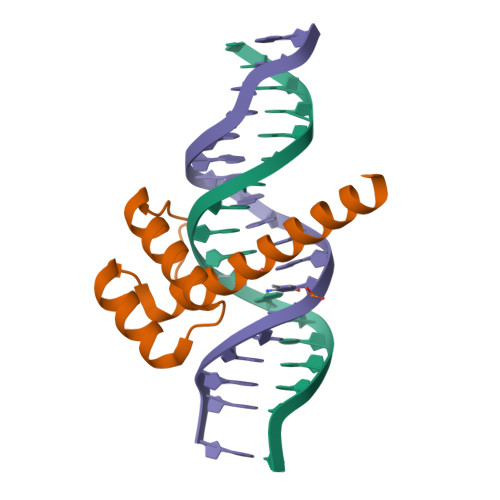
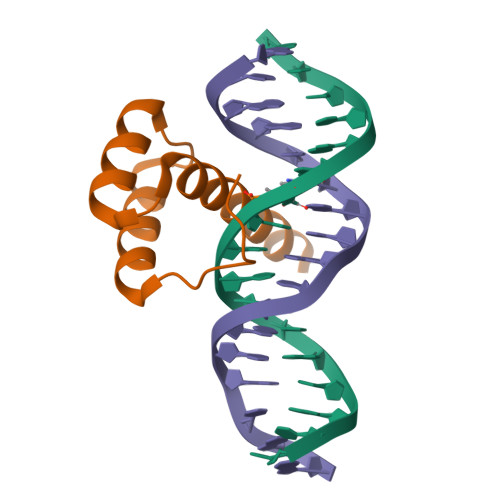
Structure of LHX4 transcription factor complexed with DNAMorgunova, E., Yin, Y., Popov, A., Taipale, J. (2017) Science 356:
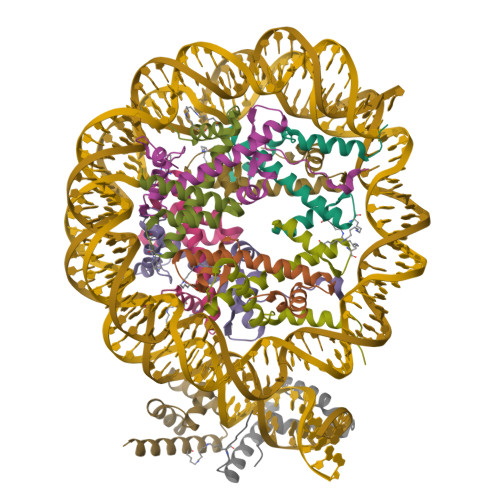
Homeobox transcription factor CDX2 bound to methylated DNAMorgunova, E., Popov, A., Taipale, J. (2017) Science 356:
Homeobox transcription factor CDX1 bound to methylated DNAMorgunova, E., Popov, A., Taipale, J. (2017) Science 356:
Cryo-EM structure of NCP-6-4PP(-1)-UV-DDBMatsumoto, S., Cavadini, S., Bunker, R.D., Thoma, N.H. (2019) Nature 571: 79-84
Cryo-EM structure of NCP_THF2(-1)-UV-DDBMatsumoto, S., Cavadini, S., Bunker, R.D., Thoma, N.H. (2019) Nature 571: 79-84
Cryo-EM structure of NCP-THF2(+1)-UV-DDB class AMatsumoto, S., Cavadini, S., Bunker, R.D., Thoma, N.H. (2019) Nature 571: 79-84
Cryo-EM structure of NCP_THF2(-3)-UV-DDBMatsumoto, S., Cavadini, S., Bunker, R.D., Thoma, N.H. (2019) Nature 571: 79-84
Cryo-EM structure of NCP-THF2(+1)-UV-DDB class BMatsumoto, S., Cavadini, S., Bunker, R.D., Thoma, N.H. (2019) Nature 571: 79-84
Cryo-EM structure of NCP-6-4PPMatsumoto, S., Cavadini, S., Bunker, R.D., Thoma, N.H. (2019) Nature 571: 79-84
Cryo-EM structure of NCP_THF2(-3)Matsumoto, S., Cavadini, S., Bunker, R.D., Thoma, N.H. (2019) Nature 571: 79-84
OCT4-SOX2-bound nucleosome - SHL-6Michael, A.K., Kempf, G., Cavadini, S., Bunker, R.D., Thoma, N.H. (2020) Science 368: 1460-1465
Nucleosome with OCT4-SOX2 motif at SHL-6Michael, A.K., Kempf, G., Cavadini, S., Bunker, R.D., Thoma, N.H. (2020) Science 368: 1460-1465
OCT4-SOX2-bound nucleosome - SHL+6Michael, A.K., Kempf, G., Cavadini, S., Bunker, R.D., Thoma, N.H. (2020) Science 368: 1460-1465
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL-6.2 (DNA conformation 1)Michael, A.K., Stoos, L., Kempf, G., Cavadini, S., Thoma, N.H. (2023) Nature 619: 385-393
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (composite map)Stoos, L., Michael, A.K., Kempf, G., Cavadini, S., Thoma, N.H. (2023) Nature 619: 385-393
Cryo-EM structure of CLOCK-BMAL1 bound to the native Por enhancer nucleosome (map 2, additional 3D classification and flexible refinement)Michael, A.K., Stoos, L., Kempf, G., Cavadini, S., Thoma, N. (2023) Nature 619: 385-393
OCT4 and MYC-MAX co-bound to a nucleosomeMichael, A.K., Stoos, L., Kempf, G., Cavadini, S., Thoma, N. (2023) Nature 619: 385-393
MYC-MAX bound to a nucleosome at SHL+5.8Stoos, L., Michael, A.K., Kempf, G., Kater, L., Cavadini, S., Thoma, N. (2023) Nature 619: 385-393
1 to 20 of 20 Structures Page 1 of 1 Sort by |