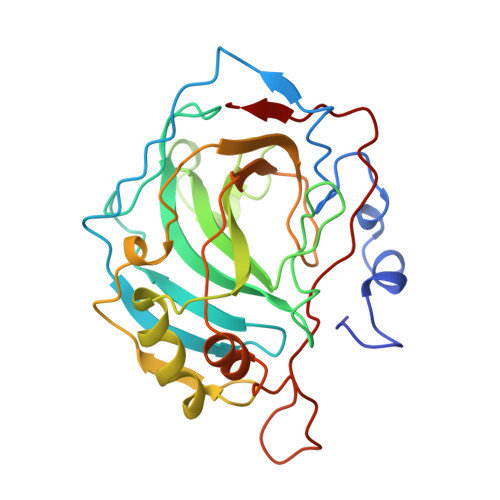
Structure of an engineered His3Cys zinc binding site in human carbonic anhydrase II.
Ippolito, J.A., Christianson, D.W.(1993) Biochemistry 32: 9901-9905
- PubMed: 8399159 Search on PubMed
- DOI: https://doi.org/10.1021/bi00089a005
- Primary Citation Related Structures:
1DCA, 1DCB - PubMed Abstract:
X-ray crystallographic analysis of the Thr-199-->Cys (T199C) variant of human carbonic anhydrase II reveals the first high-resolution structure of an engineered zinc coordination polyhedron in a metalloenzyme. In the wild-type enzyme, Thr-199 accepts a hydrogen bond from zinc-bound hydroxide; in the variant, the polypeptide backbone is sufficiently plastic to permit Cys-199 to displace hydroxide ion and coordinate to zinc with nearly perfect coordination stereochemistry. Importantly, the resulting His3-Cys-Zn2+ motif binds zinc more tightly than the wild-type enzyme [Kiefer, L. L., Krebs, J. F., Paterno, S. A., & Fierke C. A. (1993) Biochemistry (preceding paper in this issue)]. This novel zinc coordination polyhedron is analogous to that postulated for matrix metalloproteinase zymogens such as prostromelysin, where a cysteine-zinc interaction is responsible for the inactivity of the zymogen. Intriguingly, Cys-199 of T199C CAII is displaced from zinc coordination by soaking crystals in high concentrations of acetazolamide. Hence, residual catalytic activity measured for this variant probably arises from an alternate conformer of Cys-199 which allows the catalytic nucleophile, hydroxide ion, to be activated by zinc coordination.
- Department of Chemistry, University of Pennsylvania, Philadelphia 19104-6323.
Organizational Affiliation: