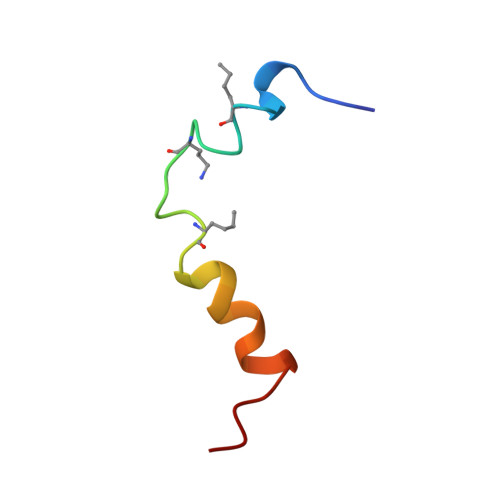
The structure of human parathyroid hormone-related protein(1-34) in near-physiological solution.
Weidler, M., Marx, U.C., Seidel, G., Schaefer, W., Hoffmann, E., Esswein, A., Roesch, P.(1999) FEBS Lett 444: 239-244
- PubMed: 10050767 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(98)01658-5
- Primary Citation Related Structures:
1BZG, 1HTH - PubMed Abstract:
Parathyroid hormone-related protein plays a major role in the pathogenesis of humoral hypercalcemia of malignancy. Under normal physiological conditions, parathyroid hormone-related protein is produced in a wide variety of tissues and acts in an autocrine or paracrine fashion. Parathyroid hormone-related protein and parathyroid hormone bind to and activate the same G-protein-coupled receptor. Here we present the structure of the biologically active NH2-terminal domain of human parathyroid hormone-related protein(1-34) in near-physiological solution in the absence of crowding reagents as determined by two-dimensional proton magnetic resonance spectroscopy. An improved strategy for structure calculation revealed the presence of two helices, His-5-Leu-8 and Gln-16-Leu-27, connected by a flexible linker. The parathyroid hormone-related protein(1-34) structure and the structure of human parathyroid hormone(1-37) as well as human parathyroid hormone(1-34) are highly similar, except for the well defined turn, His-14-Ser-17, present in parathyroid hormone. Thus, the similarity of the binding affinities of parathyroid hormone and parathyroid hormone-related protein to their common receptor may be based on their structural similarity.
- Lehrstuhl für Biopolymere, Universität Bayreuth, Germany.
Organizational Affiliation: