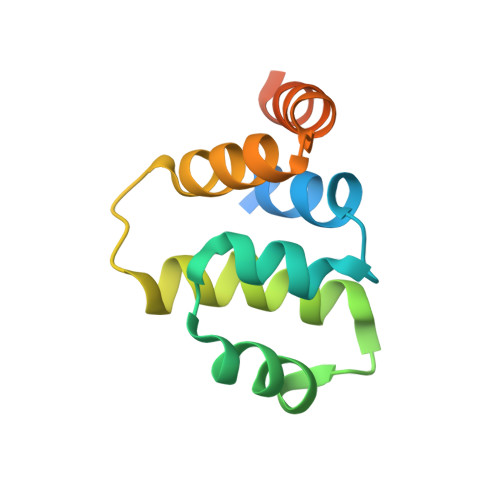
Solution structure of the tumor necrosis factor receptor-1 death domain.
Sukits, S.F., Lin, L.L., Hsu, S., Malakian, K., Powers, R., Xu, G.Y.(2001) J Mol Biology 310: 895-906
- PubMed: 11453696
- DOI: https://doi.org/10.1006/jmbi.2001.4790
- Primary Citation of Related Structures:
1ICH - PubMed Abstract:
Tumor necrosis factor receptor-1 death domain (TNFR-1 DD) is the intracellular functional domain responsible for the receptor signaling activities. The solution structure of the R347K mutant of TNFR-1 DD was solved by NMR spectroscopy. A total of 20 structures were calculated by means of hybrid distance geometry-simulated annealing using a total of 1167 distance constraints and 117 torsion angle constraints. The atomic rms distribution about the mean coordinate positions for the 20 structures for residues composing the secondary structure region is 0.40 A for the backbone atoms and 1.09 A for all atoms. The structure consists of six antiparallel alpha-helices arranged in a similar fashion to the other members of the death domain superfamily. The secondary structure and three-dimensional structure of R347K TNFR1-DD are very similar to the secondary structure and deduced topology of the R347A TNFR1-DD mutant. Mutagenesis studies identified critical residues located in alpha2 and part of alpha3 and alpha4 that are crucial for self-interaction and interaction with TRADD. Structural superposition with previously solved proteins in the death domain superfamily reveals that the major differences between the structures reside in alpha2, alpha3, and alpha4. Interestingly, these regions correspond to the binding sites of TNFR1-DD, providing a structural basis for the specificity of death domain interactions and its subsequent signaling event.
- Department of Biological Chemistry, Wyeth Research, Cambridge, MA 02140, USA.
Organizational Affiliation: