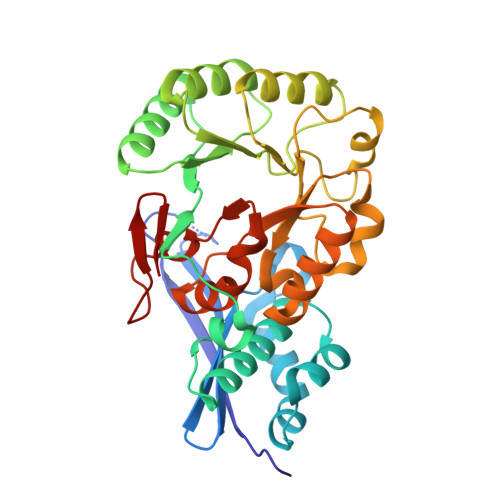
Evolution of enzymatic activities in the enolase superfamily: crystal structures of the L-Ala-D/L-Glu epimerases from Escherichia coli and Bacillus subtilis.
Gulick, A.M., Schmidt, D.M., Gerlt, J.A., Rayment, I.(2001) Biochemistry 40: 15716-15724
- PubMed: 11747448 Search on PubMed
- DOI: https://doi.org/10.1021/bi011641p
- Primary Citation Related Structures:
1JPD, 1JPM - PubMed Abstract:
The members of the enolase superfamily catalyze different overall reactions, yet share a partial reaction that involves Mg(2+)-assisted enolization of the substrate carboxylate anion. The fate of the resulting enolate intermediate is determined by the active site of each enzyme. Several members of this superfamily have been structurally characterized to permit an understanding of the evolutionary strategy for using a common structural motif to catalyze different overall reactions. In the preceding paper, two new members of the superfamily were identified that catalyze the epimerization of the glutamate residue in L-Ala-D/L-Glu. These enzymes belong to the muconate lactonizing enzyme subgroup of the enolase superfamily, and their sequences are only 31% identical. The structure of YcjG, the epimerase from Escherichia coli, was determined by MAD phasing using both the SeMet-labeled protein and a heavy atom derivative. The structure of YkfB, the epimerase from Bacillus subtilis, was determined by molecular replacement using the muconate lactonizing enzyme as a search model. In this paper, we report the three-dimensional structures of these enzymes and compare them to the structure of o-succinylbenzoate synthase, another member of the muconate lactonizing enzyme subgroup.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: