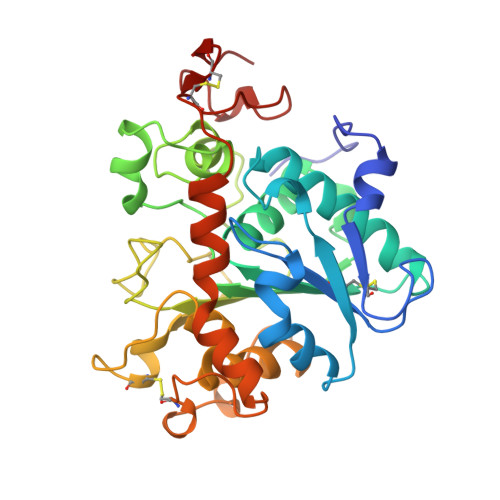
Crystallographic and molecular-modeling studies of lipase B from Candida antarctica reveal a stereospecificity pocket for secondary alcohols.
Uppenberg, J., Ohrner, N., Norin, M., Hult, K., Kleywegt, G.J., Patkar, S., Waagen, V., Anthonsen, T., Jones, T.A.(1995) Biochemistry 34: 16838-16851
- PubMed: 8527460
- DOI: https://doi.org/10.1021/bi00051a035
- Primary Citation Related Structures:
1LBS, 1LBT - PubMed Abstract:
Many lipases are potent catalysts of stereoselective reactions and are therefore of interest for use in chemical synthesis. The crystal structures of lipases show a large variation in the shapes of their active site environments that may explain the large variation in substrate specificity of these enzymes. We have determined the three-dimensional structure of Candida antarctica lipase B (CALB) cocrystallized with the detergent Tween 80. In another crystal form, the structure of the enzyme in complex with a covalently bound phosphonate inhibitor has been determined. In both structures, the active site is exposed to the external solvent. The potential lid-forming helix alpha 5 in CALB is well-ordered in the Tween 80 structure and disordered in the inhibitor complex. The tetrahedral intermediates of two chiral substrates have been modeled on the basis of available structural and biochemical information. The results of this study provide a structural explanation for the high stereoselectivity of CALB toward many secondary alcohols.
- Department of Molecular Biology, Uppsala University, Sweden.
Organizational Affiliation: