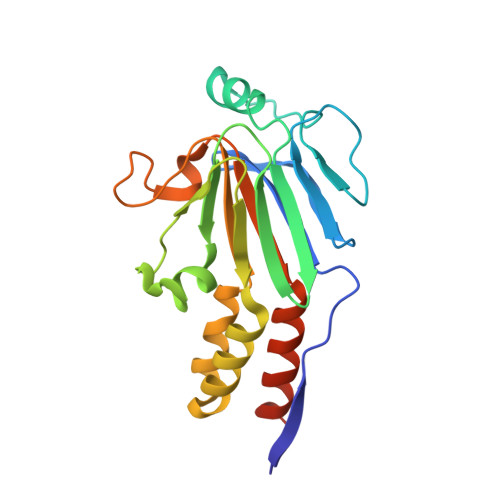
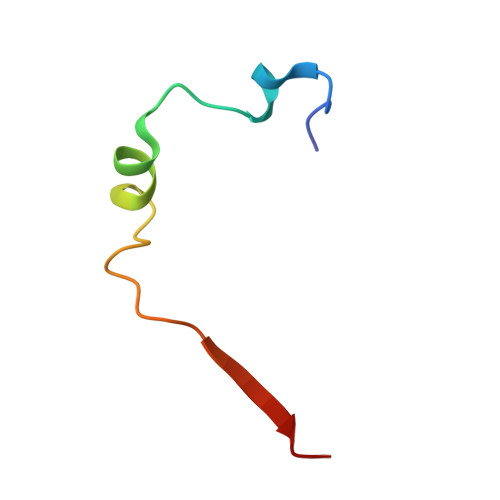
Smad3 allostery links TGF-beta receptor kinase activation to transcriptional control
Qin, B.Y., Lam, S.S., Correia, J.J., Lin, K.(2002) Genes Dev 16: 1950-1963
- PubMed: 12154125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.1002002
- Primary Citation Related Structures:
1MJS, 1MK2 - PubMed Abstract:
Smad3 transduces the signals of TGF-betas, coupling transmembrane receptor kinase activation to transcriptional control. The membrane-associated molecule SARA (Smad Anchor for Receptor Activation) recruits Smad3 for phosphorylation by the receptor kinase. Upon phosphorylation, Smad3 dissociates from SARA and enters the nucleus, in which its transcriptional activity can be repressed by Ski. Here, we show that SARA and Ski recognize specifically the monomeric and trimeric forms of Smad3, respectively. Thus, trimerization of Smad3, induced by phosphorylation, simultaneously activates the TGF-beta signal by driving Smad3 dissociation from SARA and sets up the negative feedback mechanism by Ski. Structural models of the Smad3/SARA/receptor kinase complex and Smad3/Ski complex provide insights into the molecular basis of regulation.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, Massachusetts 01655, USA.
Organizational Affiliation: