Impact of Fc Glycans on The Effector Functions Vary with the Antibody mechanism of Action
Raju, T.S., Mulkerrin, M.G., Parker, M., De Vos, A.M., Gazzano-Santoro, H., Totpal, K., Ultsch, M.H.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| immunoglobulin gamma-1 heavy chain constant region | A, C [auth B] | 212 | Homo sapiens | Mutation(s): 0 | 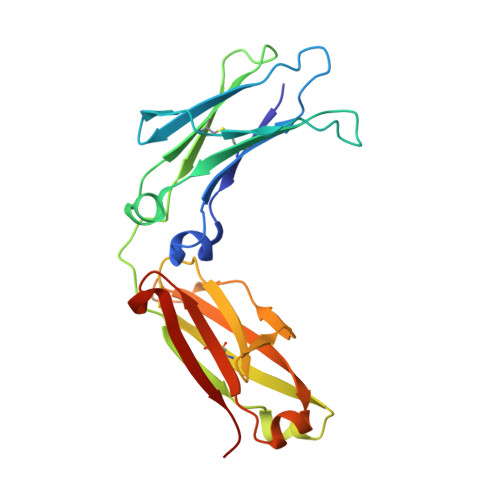 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P01857 GTEx: ENSG00000211896 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P01857 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P01857-2 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Minimized version of Protein A (Z34C) | B [auth C], D | 34 | synthetic construct | Mutation(s): 0 | 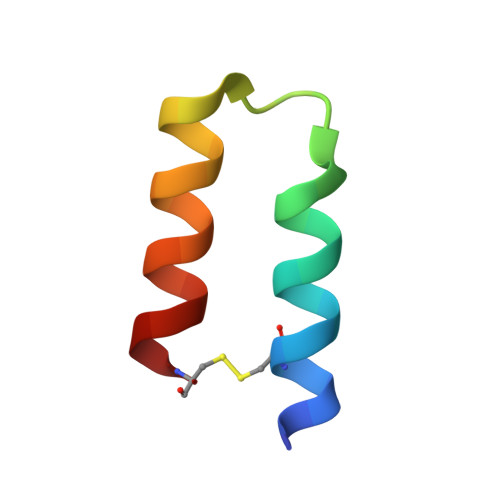 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[beta-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | E | 8 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G59451NL GlyCosmos: G59451NL GlyGen: G59451NL | |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | F | 8 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G50842MS GlyCosmos: G50842MS GlyGen: G50842MS | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 85.782 | α = 90 |
| b = 126.413 | β = 90 |
| c = 54.743 | γ = 90 |
| Software Name | Purpose |
|---|---|
| X-PLOR | refinement |
| DENZO | data reduction |
| SCALEPACK | data scaling |
| AMoRE | phasing |