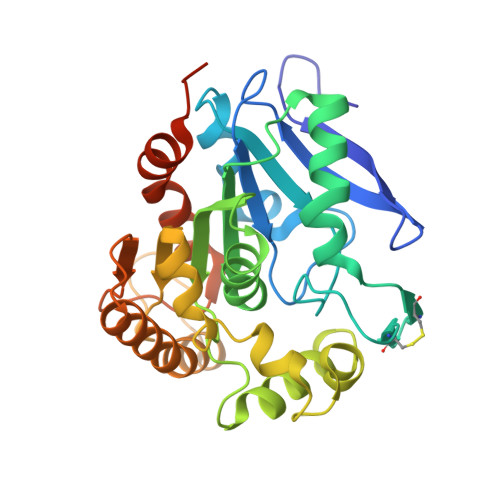
Mycobacterium tuberculosis Antigen 85A and 85C Structures Confirm Binding Orientation and Conserved Substrate Specificity
Ronning, D.R., Vissa, V., Besra, G.S., Belisle, J.T., Sacchettini, J.C.(2004) J Biol Chem 279: 36771-36777
- PubMed: 15192106
- DOI: https://doi.org/10.1074/jbc.M400811200
- Primary Citation of Related Structures:
1SFR, 1VA5 - PubMed Abstract:
The maintenance of the highly hydrophobic cell wall is central to the survival of Mycobacterium tuberculosis within its host environment. The antigen 85 proteins (85A, 85B, and 85C) of M. tuberculosis help maintain the integrity of the cell wall 1) by catalyzing the transfer of mycolic acids to the cell wall arabinogalactan and 2) through the synthesis of trehalose dimycolate (cord factor). Additionally, these secreted proteins allow for rapid invasion of alveolar macrophages via direct interactions between the host immune system and the invading bacillus. Here we describe two crystal structures: the structure of antigen 85C co-crystallized with octylthioglucoside as substrate, resolved to 2.0 A, and the crystal structure of antigen 85A, which was solved at a resolution of 2.7 A. The structure of 85C with the substrate analog identifies residues directly involved in substrate binding. Elucidation of the antigen 85A structure, the last of the three antigen 85 homologs to be solved, shows that the active sites of the three antigen 85 proteins are virtually identical, indicating that these share the same substrate. However, in contrast to the high level of conservation within the substrate-binding site and the active site, surface residues disparate from the active site are quite variable, indicating that three antigen 85 enzymes are needed to evade the host immune system.
Organizational Affiliation:
Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77843-2128, USA.