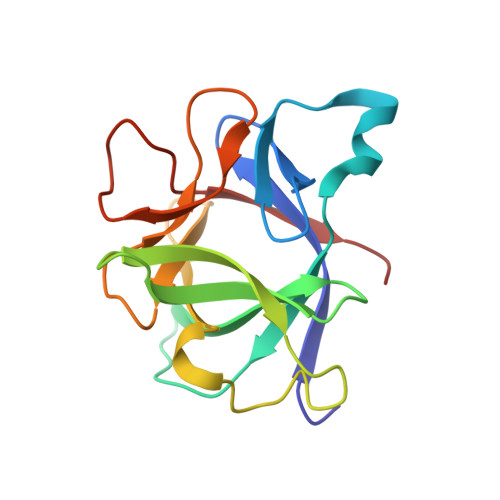
Structural and energetic consequences of mutations in a solvated hydrophobic cavity.
Adamek, D.H., Guerrero, L., Blaber, M., Caspar, D.L.(2005) J Mol Biology 346: 307-318
- PubMed: 15663946 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.11.046
- Primary Citation Related Structures:
1S0L, 1T4Q, 1TOO, 1TP0, 1TWE, 1TWM - PubMed Abstract:
The structural and energetic consequences of modifications to the hydrophobic cavity of interleukin 1-beta (IL-1beta) are described. Previous reports demonstrated that the entirely hydrophobic cavity of IL-1beta contains positionally disordered water. To gain a better understanding of the nature of this cavity and the water therein, a number of mutant proteins were constructed by site-directed mutagenesis, designed to result in altered hydrophobicity of the cavity. These mutations involve the replacement of specific phenylalanine residues, which circumscribe the cavity, with tyrosine, tryptophan, leucine and isoleucine. Using differential scanning calorimetry to determine the relative stabilities of the wild-type and mutant proteins, we found all of the mutants to be destabilizing. X-ray crystallography was used to identify the structural consequences of the mutations. No clear correlation between the hydrophobicities of the specific side-chains introduced and the resulting stabilities was found.
- Institute of Molecular Biophysics, Florida State University, Tallahassee, FL 32306-4380, USA. daniel.adamek@msfc.nasa.gov
Organizational Affiliation: