Synthesis and conformational analysis of a non-amidine factor Xa inhibitor that incorporates 5-methyl-4,5,6,7-tetrahydrothiazolo[5,4-c]pyridine as S4 binding element
Haginoya, N., Kobayashi, S., Komoriya, S., Yoshino, T., Suzuki, M., Shimada, T., Watanabe, K., Hirokawa, Y., Furugori, T., Nagahara, T.(2004) J Med Chem 47: 5167-5182
- PubMed: 15456260 Search on PubMed
- DOI: https://doi.org/10.1021/jm049884d
- Primary Citation Related Structures:
1V3X - PubMed Abstract:
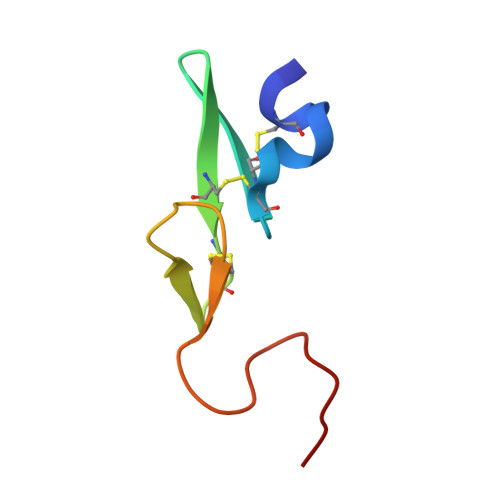
Our exploratory study was based on the concept that a non-amidine factor Xa (fXa) inhibitor is suitable for an orally available anticoagulant. We synthesized and evaluated a series of N-(6-chloronaphthalen-2-yl)sulfonylpiperazine derivatives incorporating various fused-bicyclic rings containing an aliphatic amine expected to be S4 binding element. Among this series, 5-methyl-4,5,6,7-tetrahydrothiazolo[5,4-c]pyridine type 61 displayed orally potent anti-fXa activity and evident prolongation of prothrombin time (PT) with the moderate bioavailability in rats. The X-ray crystal analysis afforded an obvious binding mode that 5-methyl-4,5,6,7-tetrahydrothiazolo[5,4-c]pyridine and 6-chloronaphthalene respectively bound to S4 and S1 subsites. In this X-ray study, we discovered a novel intramolecular S-O close contact. Ab initio energy calculations of model compounds deduced that conformers with the most close S-O proximity were most stable. The Mulliken population analysis proposed that this energy profile was caused by both of electrostatic S-O affinity and N-O repulsion. The results of these calculations and X-ray analysis suggested a possibility that the restricted conformation effected the affinity to S4 subsite of fXa.
- Medicinal Chemistry Research Laboratory, Discovery Research Laboratory, Drug Metabolism & Physicochemical Property Research Laboratory, Daiichi Pharmaceutical Co. Ltd, 1-16-13, Kita-Kasai, Edogawa-ku, Tokyo 134-8630, Japan. hagin9kg@daiichipharm.co.jp
Organizational Affiliation: