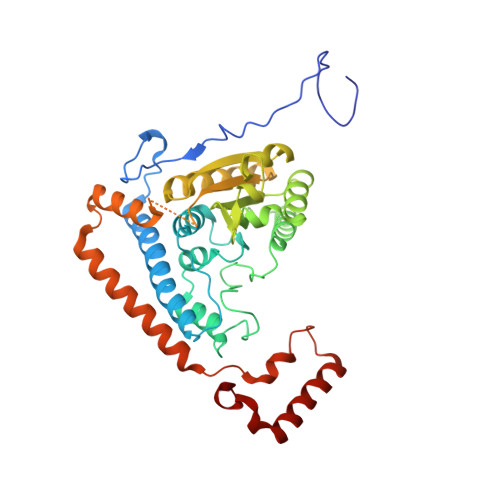
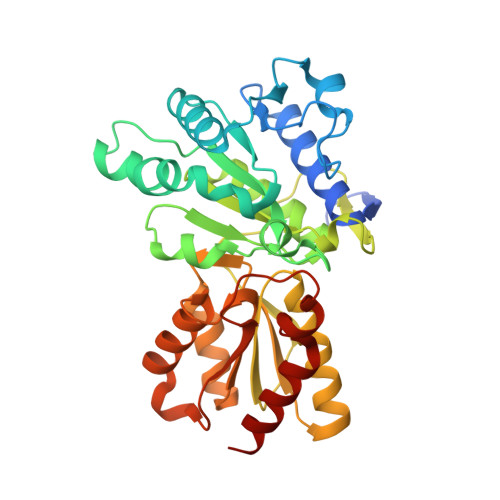
Molecular mechanism for regulation of the human mitochondrial branched-chain alpha-ketoacid dehydrogenase complex by phosphorylation
Wynn, R.M., Kato, M., Machius, M., Chuang, J.L., Li, J., Tomchick, D.R., Chuang, D.T.(2004) Structure 12: 2185-2196
- PubMed: 15576032 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.09.013
- Primary Citation Related Structures:
1U5B, 1X7W, 1X7X, 1X7Y, 1X7Z, 1X80 - PubMed Abstract:
The human mitochondrial branched-chain alpha-ketoacid dehydrogenase complex (BCKDC) is a 4 MDa macromolecular machine comprising three catalytic components (E1b, E2b, and E3), a kinase, and a phosphatase. The BCKDC overall activity is tightly regulated by phosphorylation in response to hormonal and dietary stimuli. We report that phosphorylation of Ser292-alpha in the E1b active site channel results in an order-to-disorder transition of the conserved phosphorylation loop carrying the phosphoryl serine. The conformational change is triggered by steric clashes of the phosphoryl group with invariant His291-alpha that serves as an indispensable anchor for the phosphorylation loop through bound thiamin diphosphate. Phosphorylation of Ser292-alpha does not severely impede the E1b-dependent decarboxylation of alpha-ketoacids. However, the disordered loop conformation prevents phosphorylated E1b from binding the E2b lipoyl-bearing domain, which effectively shuts off the E1b-catalyzed reductive acylation reaction and therefore completely inactivates BCKDC. This mechanism provides a paradigm for regulation of mitochondrial alpha-ketoacid dehydrogenase complexes by phosphorylation.
- Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: