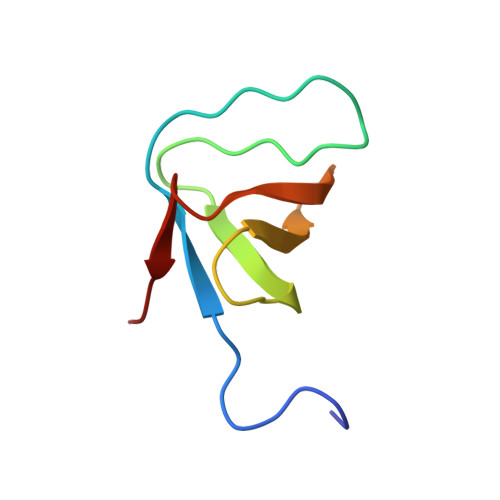
Structural Analysis of the SH3 Domain of beta-PIX and Its Interaction with alpha-p21 Activated Kinase (PAK)
Mott, H.R., Nietlispach, D., Evetts, K.A., Owen, D.(2005) Biochemistry 44: 10977-10983
- PubMed: 16101281 Search on PubMed
- DOI: https://doi.org/10.1021/bi050374a
- Primary Citation Related Structures:
1ZSG - PubMed Abstract:
The PAK Ser/Thr kinases are important downstream effectors of the Rho family GTPases Cdc42 and Rac, partly mediating the role of these G proteins in cell proliferation and cytoskeletal rearrangements. As well as small G proteins, PAK interacts with the Cdc42/Rac exchange factor beta-PIX via the PIX SH3 domain and a nontypical Pro-rich region in PAK. This interaction is thought to affect the localization of PAK, as well as increased GTP/GDP exchange of Rac and Cdc42. We have determined the structure of the PIX-SH3/PAK peptide complex and shown that it differs from typical Src-like SH3/peptide complexes. The peptide makes contacts through the Pro-rich sequence in a similar way to standard SH3/peptide complexes, even though the Pro residue positions are not conserved. In addition, there are interactions with a Pro and Lys in the PAK, which are C-terminal to the conserved Arg found in all SH3-binding sequences. These contact a fourth binding pocket on the SH3 domain. We have measured the affinity of PIX-SH3 for the PAK peptide and found that it is of intermediate affinity. When PAK is activated, Ser-199 in the PIX-binding site is phosphorylated. This phosphorylation is sufficient to reduce the affinity for PIX 6-fold.
- Department of Biochemistry, University of Cambridge, UK. mott@bioc.cam.ac.uk
Organizational Affiliation: