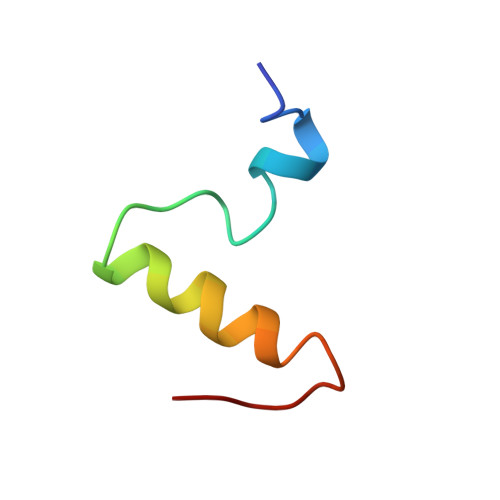
Structure-activity relation of NH2-terminal human parathyroid hormone fragments.
Marx, U.C., Adermann, K., Bayer, P., Meyer, M., Forssmann, W.G., Rosch, P.(1998) J Biological Chem 273: 4308-4316
- PubMed: 9468478
- DOI: https://doi.org/10.1074/jbc.273.8.4308
- Primary Citation Related Structures:
1ZWB, 1ZWD, 1ZWE - PubMed Abstract:
Human parathyroid hormone (hPTH) is involved in the regulation of the calcium level in blood. This hormone function is located in the NH2-terminal 34 amino acids of the 84-amino acid peptide hormone and is transduced via the adenylate cyclase and the phosphatidylinositol signaling pathways. It is well known that truncation of the two NH2-terminal amino acids of the hormone leads to complete loss of in vivo normocalcemic function. To correlate loss of calcium level regulatory activity after stepwise NH2-terminal truncation and solution structure, we studied the conformations of fragments hPTH-(2-37), hPTH-(3-37), and hPTH-(4-37) in comparison to hPTH-(1-37) in aqueous buffer solution under near physiological conditions by circular dichroism spectroscopy, two-dimensional nuclear magnetic resonance spectroscopy, and restrained molecular dynamics calculations. All peptides show helical structures and hydrophobic interactions between Leu-15 and Trp-23 that lead to a defined loop region from His-14 to Ser-17. A COOH-terminal helix from Met-18 to at least Leu-28 was found for all peptides. The helical structure in the NH2-terminal part of the peptides was lost in parallel with the NH2-terminal truncation and can be correlated with the loss of calcium regulatory activity.
- Lehrstuhl für Biopolymere, Universität Bayreuth, D-95440 Bayreuth, Federal Republic of Germany.
Organizational Affiliation: