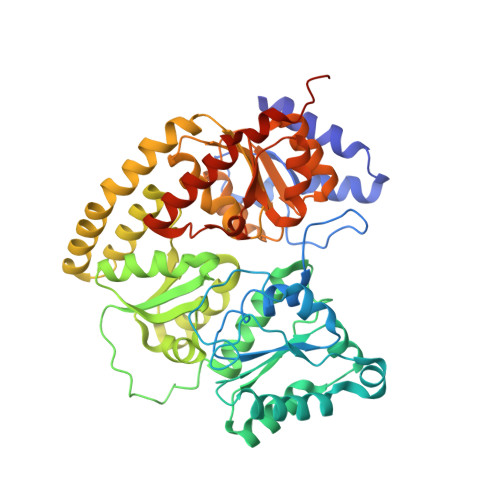
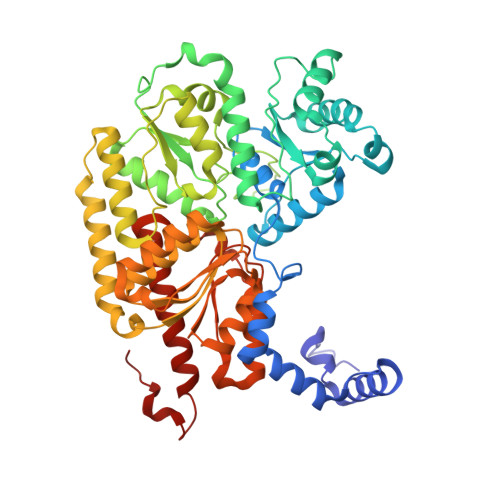
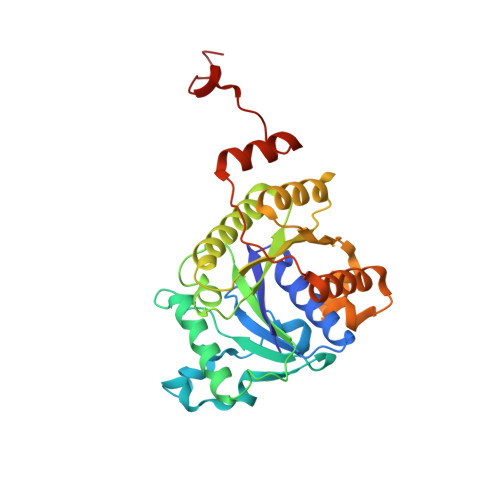
Nitrogenase Complexes: Multiple Docking Sites for a Nucleotide Switch Protein
Tezcan, F.A., Kaiser, J.T., Mustafi, D., Walton, M.Y., Howard, J.B., Rees, D.C.(2005) Science 309: 1377-1380
- PubMed: 16123301 Search on PubMed
- DOI: https://doi.org/10.1126/science.1115653
- Primary Citation Related Structures:
2AFH, 2AFI, 4WZB - PubMed Abstract:
Adenosine triphosphate (ATP) hydrolysis in the nitrogenase complex controls the cycle of association and dissociation between the electron donor adenosine triphosphatase (ATPase) (Fe-protein) and its target catalytic protein (MoFe-protein), driving the reduction of dinitrogen into ammonia. Crystal structures in different nucleotide states have been determined that identify conformational changes in the nitrogenase complex during ATP turnover. These structures reveal distinct and mutually exclusive interaction sites on the MoFe-protein surface that are selectively populated, depending on the Fe-protein nucleotide state. A consequence of these different docking geometries is that the distance between redox cofactors, a critical determinant of the intermolecular electron transfer rate, is coupled to the nucleotide state. More generally, stabilization of distinct docking geometries by different nucleotide states, as seen for nitrogenase, could enable nucleotide hydrolysis to drive the relative motion of protein partners in molecular motors and other systems.
- Division of Chemistry and Chemical Engineering, California Institute of Technology, Mail Code 114-96, Pasadena, CA 91125, USA.
Organizational Affiliation: