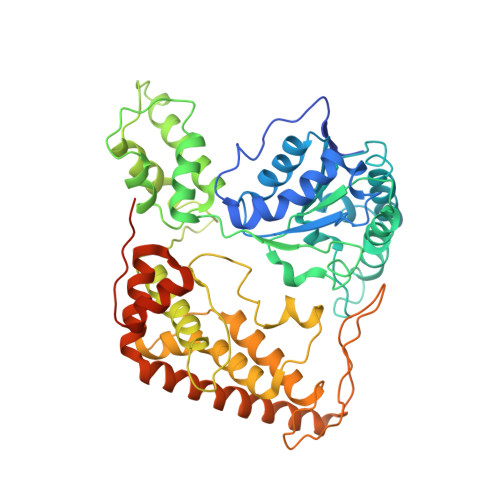
Structure of the Whole Cytosolic Region of ATP-Dependent Protease FtsH
Suno, R., Niwa, H., Tsuchiya, D., Zhang, X., Yoshida, M., Morikawa, K.(2006) Mol Cell 22: 575-585
- PubMed: 16762831 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2006.04.020
- Primary Citation Related Structures:
2DHR, 2DI4, 4EIW - PubMed Abstract:
An ATP-dependent protease, FtsH, digests misassembled membrane proteins in order to maintain membrane integrity and digests short-lived soluble proteins in order to control their cellular regulation. This enzyme has an N-terminal transmembrane segment and a C-terminal cytosolic region consisting of an AAA+ ATPase domain and a protease domain. Here we present two crystal structures: the protease domain and the whole cytosolic region. The cytosolic region fully retains an ATP-dependent protease activity and adopts a three-fold-symmetric hexameric structure. The protease domains displayed a six-fold symmetry, while the AAA+ domains, each containing ADP, alternate two orientations relative to the protease domain, making "open" and "closed" interdomain contacts. Apparently, ATPase is active only in the closed form, and protease operates in the open form. The protease catalytic sites are accessible only through a tunnel following from the AAA+ domain of the adjacent subunit, raising a possibility of translocation of polypeptide substrate to the protease sites through this tunnel.
- Chemical Resources Laboratory, Tokyo Institute of Technology, 4259 Nagatsuta, Yokohama 226-8503, Japan.
Organizational Affiliation: