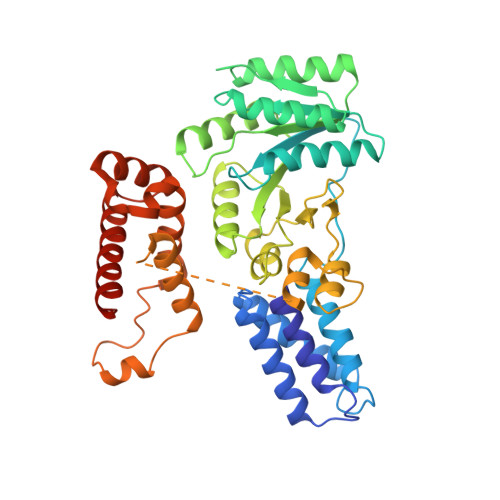
Crystal structure of the signal sequence binding subunit of the signal recognition particle.
Keenan, R.J., Freymann, D.M., Walter, P., Stroud, R.M.(1998) Cell 94: 181-191
- PubMed: 9695947 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81418-x
- Primary Citation Related Structures:
2FFH - PubMed Abstract:
The crystal structure of the signal sequence binding subunit of the signal recognition particle (SRP) from Thermus aquaticus reveals a deep groove bounded by a flexible loop and lined with side chains of conserved hydrophobic residues. The groove defines a flexible, hydrophobic environment that is likely to contribute to the structural plasticity necessary for SRP to bind signal sequences of different lengths and amino acid sequence. The structure also reveals a helix-turn-helix motif containing an arginine-rich alpha helix that is required for binding to SRP RNA and is implicated in forming the core of an extended RNA binding surface.
- Department of Biochemistry and Biophysics, School of Medicine, University of California, San Francisco 94143-0448, USA.
Organizational Affiliation: