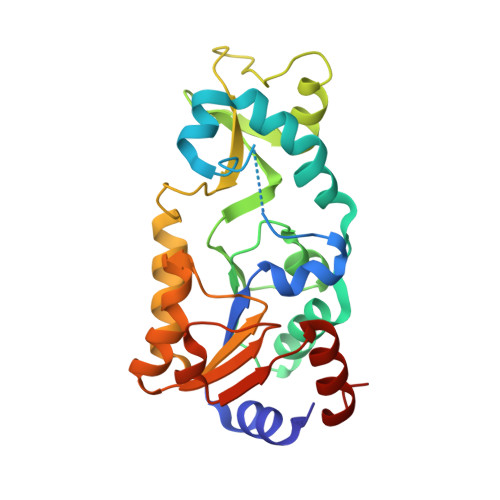
Insights into the Sirtuin Mechanism from Ternary Complexes Containing NAD(+) and Acetylated Peptide.
Hoff, K.G., Avalos, J.L., Sens, K., Wolberger, C.(2006) Structure 14: 1231-1240
- PubMed: 16905097 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2006.06.006
- Primary Citation Related Structures:
2H4F, 2H4H, 2H4J, 2H59 - PubMed Abstract:
Sirtuin proteins comprise a unique class of NAD+-dependent protein deacetylases. Although several structures of sirtuins have been determined, the mechanism by which NAD+ cleavage occurs has remained unclear. We report the structures of ternary complexes containing NAD+ and acetylated peptide bound to the bacterial sirtuin Sir2Tm and to a catalytic mutant (Sir2Tm(H116Y)). NAD+ in these structures binds in a conformation different from that seen in previous structures, exposing the alpha face of the nicotinamide ribose to the carbonyl oxygen of the acetyl lysine substrate. The NAD+ conformation is identical in both structures, suggesting that proper coenzyme orientation is not dependent on contacts with the catalytic histidine. We also present the structure of Sir2Tm(H116A) bound to deacteylated peptide and 3'-O-acetyl ADP ribose. Taken together, these structures suggest a mechanism for nicotinamide cleavage in which an invariant phenylalanine plays a central role in promoting formation of the O-alkylamidate reaction intermediate and preventing nicotinamide exchange.
- Department of Biophysics and Biophysical Chemistry, Howard Hughes Medical Institute, The Johns Hopkins University School of Medicine, 725 North Wolfe Street, Baltimore, Maryland 21205, USA.
Organizational Affiliation: