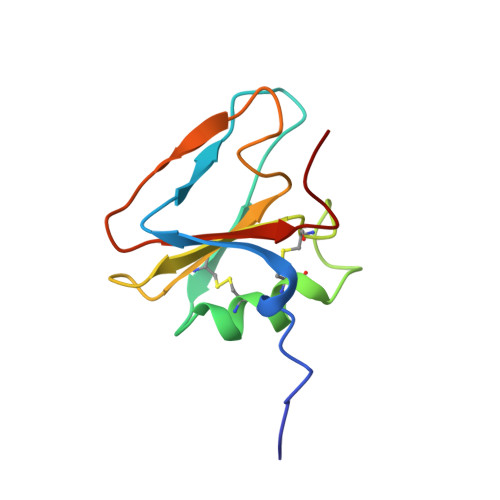
The solution structure of the N-terminal domain of hepatocyte growth factor reveals a potential heparin-binding site.
Zhou, H., Mazzulla, M.J., Kaufman, J.D., Stahl, S.J., Wingfield, P.T., Rubin, J.S., Bottaro, D.P., Byrd, R.A.(1998) Structure 6: 109-116
- PubMed: 9493272 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00012-4
- Primary Citation Related Structures:
2HGF - PubMed Abstract:
Hepatocyte growth factor (HGF) is a multipotent growth factor that transduces a wide range of biological signals, including mitogenesis, motogenesis, and morphogenesis. The N-terminal (N) domain of HGF, containing a hairpin-loop region, is important for receptor binding and the potent biological activities of HGF. The N domain is also the primary binding site for heparin or heparan sulfate, which enhances, receptor/ligand oligomerization and modulates receptor-dependent mitogenesis. The rational design of artificial modulators of HGF signaling requires a detailed understanding of the structures of HGF and its receptor, as well as the role of heparin proteoglycan; this study represents the first step towards that goal. We report here a high-resolution structure of the N domain of HGF. This first structure of HGF reveals a novel folding topology with a distinct pattern of charge distribution and indicates a possible heparin-binding site. The hairpin-loop region of the N domain plays a major role in stabilizing the structure and contributes to a putative heparin-binding site, which explains why it is required for biological functions. These results suggest several basic and/or polar residues that may be important for use in further mutational studies of heparin binding.
- Macromolecular NMR Section, ABL-Basic Research Program, NCI-Frederick Cancer Research and Development Center, Maryland 21702-1201, USA.
Organizational Affiliation: