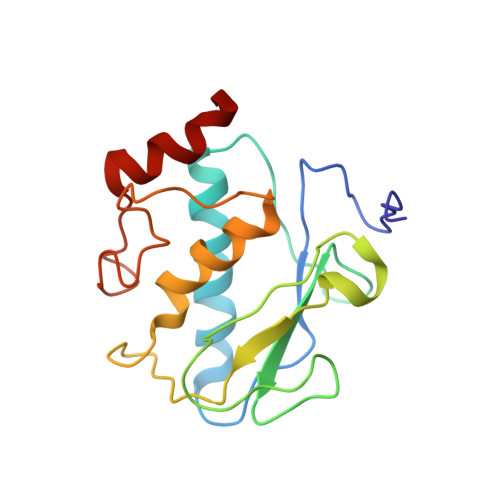
Solution structure of wild-type human matrix metalloproteinase 12 (MMP-12) in complex with a tight-binding inhibitor.
Markus, M.A., Dwyer, B., Wolfrom, S., Li, J., Li, W., Malakian, K., Wilhelm, J., Tsao, D.H.(2008) J Biomol NMR 41: 55-60
- PubMed: 18425585
- DOI: https://doi.org/10.1007/s10858-008-9236-4
- Primary Citation of Related Structures:
2K2G - Structural Biology and Computational Chemistry, Chemical and Screening Sciences, Wyeth Research, 87 Cambridge Park Drive, Cambridge, MA 02140, USA. mmarkus@wyeth.com
Organizational Affiliation: