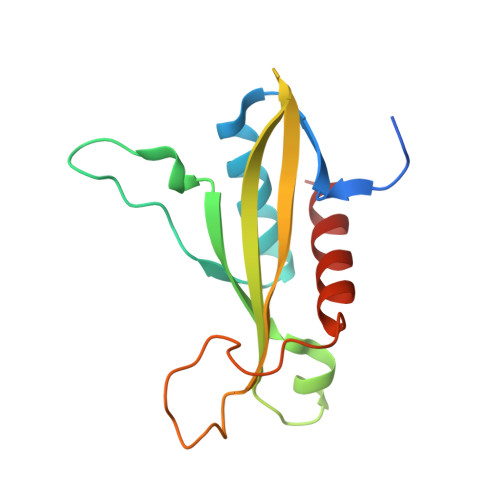
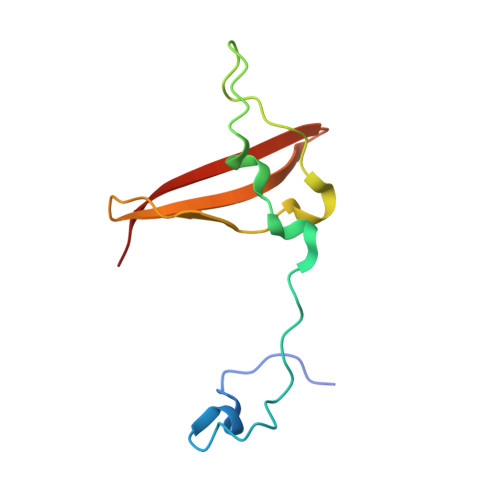
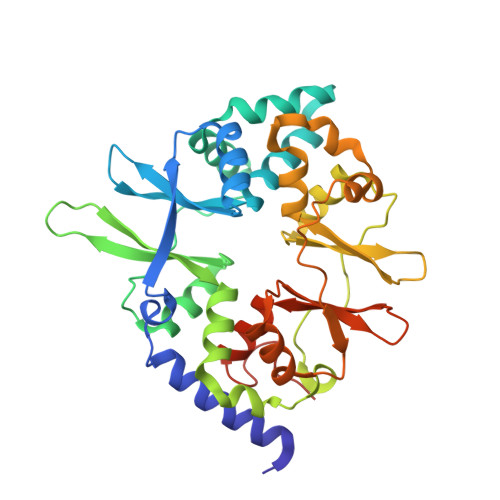
Crystal structures of the adenylate sensor from fission yeast AMP-activated protein kinase.
Townley, R., Shapiro, L.(2007) Science 315: 1726-1729
- PubMed: 17289942 Search on PubMed
- DOI: https://doi.org/10.1126/science.1137503
- Primary Citation Related Structures:
2OOX, 2OOY - PubMed Abstract:
The 5'-AMP (adenosine monophosphate)-activated protein kinase (AMPK) coordinates metabolic function with energy availability by responding to changes in intracellular ATP (adenosine triphosphate) and AMP concentrations. Here, we report crystal structures at 2.9 and 2.6 A resolution for ATP- and AMP-bound forms of a core alphabetagamma adenylate-binding domain from the fission yeast AMPK homolog. ATP and AMP bind competitively to a single site in the gamma subunit, with their respective phosphate groups positioned near function-impairing mutants. Unexpectedly, ATP binds without counterions, amplifying its electrostatic effects on a critical regulatory region where all three subunits converge.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: