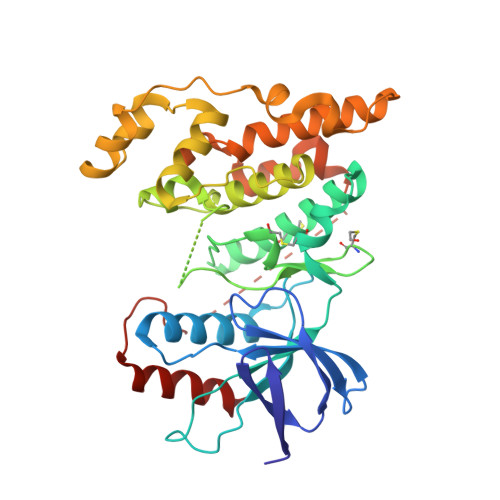
3,5-Disubstituted quinolines as novel c-Jun N-terminal kinase inhibitors.
Jiang, R., Duckett, D., Chen, W., Habel, J., Ling, Y.Y., LoGrasso, P., Kamenecka, T.M.(2007) Bioorg Med Chem Lett 17: 6378-6382
- PubMed: 17911023 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.08.054
- Primary Citation Related Structures:
2R9S - PubMed Abstract:
The structure-based design and synthesis of a novel series of c-Jun N-terminal kinase (JNK) inhibitors with selectivity against p38 is reported. The unique structure of 3,5-disubstituted quinolines (2) was developed from the previously reported 4-(2,7-phenanthrolin-9-yl)phenol (1). The X-ray crystal structure of 16a in JNK3 reveals an unexpected binding mode for this new scaffold with protein.
- Department of Medicinal Chemistry, Scripps Florida, 5353 Parkside Drive, RF-2, Jupiter, FL 33458, USA. rjiang@scripps.edu
Organizational Affiliation: