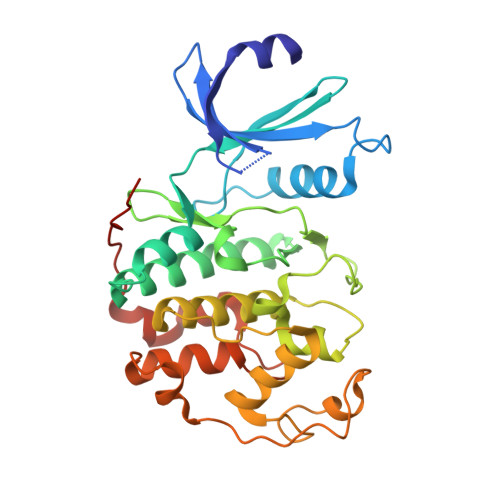
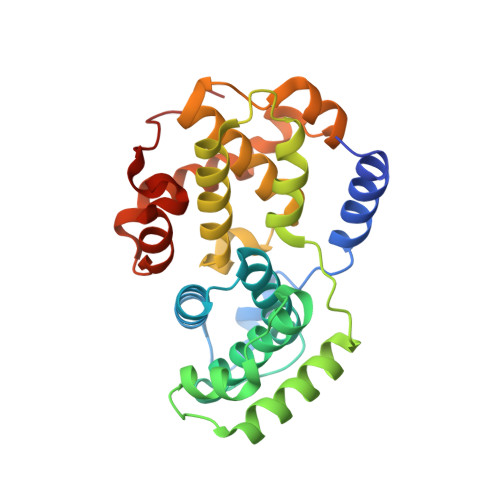
Identification of N,1,4,4-Tetramethyl-8-{[4-(4-Methylpiperazin-1-Yl)Phenyl]Amino}-4,5-Dihydro-1H-Pyrazolo[4,3-H]Quinazoline-3-Carboxamide (Pha-848125), a Potent, Orally Available Cyclin Dependent Kinase Inhibitor.
Brasca, M.G., Amboldi, N., Ballinari, D., Cameron, A., Casale, E., Cervi, G., Colombo, M., Colotta, F., Croci, V., D'Alessio, R., Fiorentini, F., Isacchi, A., Mercurio, C., Moretti, W., Panzeri, A., Pastori, W., Pevarello, P., Quartieri, F., Roletto, F., Traquandi, G., Vianello, P., Vulpetti, A., Ciomei, M.(2009) J Med Chem 52: 5152
- PubMed: 19603809 Search on PubMed
- DOI: https://doi.org/10.1021/jm9006559
- Primary Citation Related Structures:
2WIH, 2WIP - PubMed Abstract:
The discovery of a novel class of inhibitors of cyclin dependent kinases (CDKs) is described. Starting from compound 1, showing good potency as inhibitor of CDKs but being poorly selective against a panel of serine-threonine and tyrosine kinases, new analogues were synthesized. Enhancement in selectivity, antiproliferative activity against A2780 human ovarian carcinoma cells, and optimization of the physical properties and pharmacokinetic profile led to the identification of highly potent and orally available compounds. Compound 28 (PHA-848125), which in the preclinical xenograft A2780 human ovarian carcinoma model showed good efficacy and was well tolerated upon repeated daily treatments, was identified as a drug candidate for further development. Compound 28 is currently undergoing phase I and phase II clinical trials.
- Business Unit Oncology, Nerviano Medical Sciences Srl, Viale Pasteur 10, 20014 Nerviano (MI), Italy. gabriella.brasca@nervianoms.com
Organizational Affiliation: