An Extracellular Steric Seeding Mechanism for Eph-Ephrin Signalling Platform Assembly
Seiradake, E., Harlos, K., Sutton, G., Aricescu, A.R., Jones, E.Y.(2010) Nat Struct Mol Biol 17: 398
- PubMed: 20228801 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1782
- Primary Citation Related Structures:
2X10, 2X11 - PubMed Abstract:
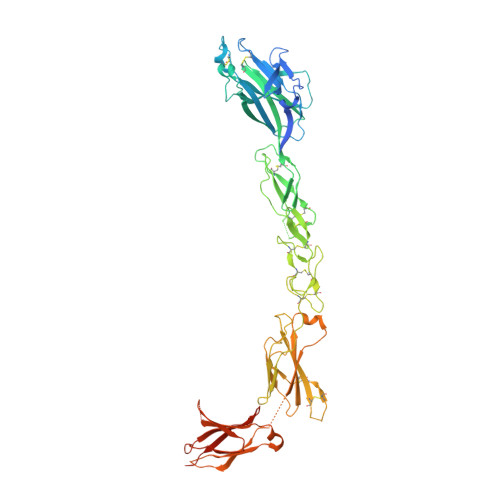
Erythropoetin-producing hepatoma (Eph) receptors are cell-surface protein tyrosine kinases mediating cell-cell communication. Upon activation, they form signaling clusters. We report crystal structures of the full ectodomain of human EphA2 (eEphA2) both alone and in complex with the receptor-binding domain of the ligand ephrinA5 (ephrinA5 RBD). Unliganded eEphA2 forms linear arrays of staggered parallel receptors involving two patches of residues conserved across A-class Ephs. eEphA2-ephrinA5 RBD forms a more elaborate assembly, whose interfaces include the same conserved regions on eEphA2, but rearranged to accommodate ephrinA5 RBD. Cell-surface expression of mutant EphA2s showed that these interfaces are critical for localization at cell-cell contacts and activation-dependent degradation. Our results suggest a 'nucleation' mechanism whereby a limited number of ligand-receptor interactions 'seed' an arrangement of receptors which can propagate into extended signaling arrays.
- Division of Structural Biology, Wellcome Trust Centre for Human Genetics, University of Oxford, Oxford, UK.
Organizational Affiliation: