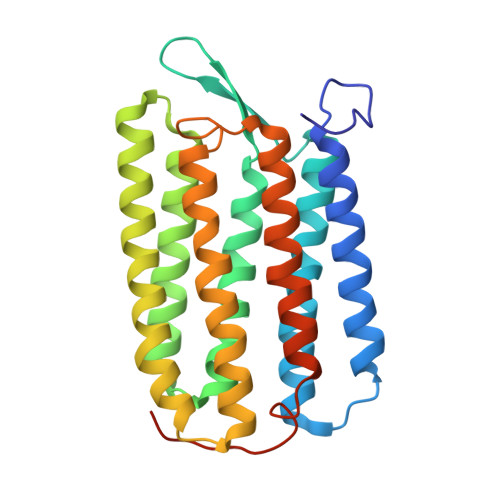
Structural role of bacterioruberin in the trimeric structure of archaerhodopsin-2
Yoshimura, K., Kouyama, T.(2008) J Mol Biology 375: 1267-1281
- PubMed: 18082767
- DOI: https://doi.org/10.1016/j.jmb.2007.11.039
- Primary Citation of Related Structures:
2EI4, 2Z55 - PubMed Abstract:
Archaerhodopsin-2 (aR2), a retinal protein-carotenoid complex found in the claret membrane of Halorubrum sp. aus-2, functions as a light-driven proton pump. In this study, the membrane fusion method was utilized to prepare trigonal P321 crystals (a=b=98.2 A, c=56.2 A) and hexagonal P6(3) crystals (a=b=108.8 A, c=220.7 A). The trigonal crystal is made up of stacked membranes in which the aR2 trimers are arranged on a honeycomb lattice. Similar membranous structures are found in the hexagonal crystal, but four membrane layers with different orientations are contained in the unit cell. In these crystals, the carotenoid bacterioruberin [5,32-bis(2-hydroxypropan-2-yl)-2,8,12,16,21,25,29,35-octamethylhexatriaconta-6,8,10,12,14,16,18,20,22,24,26,28,30-tridecaene-2,35-diol] binds to crevices between the subunits of the trimer. Its polyene chain is inclined from the membrane normal by an angle of about 20 degrees and, on the cytoplasmic side, it is surrounded by helices AB and DE of neighbouring subunits. This peculiar binding mode suggests that bacterioruberin plays a striking structural role for the trimerization of aR2. When compared with the aR2 structure in another crystal form containing no bacterioruberin, the proton release channel takes a more closed conformation in the P321 or P6(3) crystal; i.e., the native conformation of protein is stabilized in the trimeric protein-bacterioruberin complex. Interestingly, most residues participating in the trimerization are not conserved in bacteriorhodopsin, a homologous protein capable of forming a trimeric structure in the absence of bacterioruberin. Despite a large alteration in the amino acid sequence, the shape of the intratrimer hydrophobic space filled by lipids is highly conserved between aR2 and bacteriorhodopsin. Since a transmembrane helix facing this space undergoes a large conformational change during the proton pumping cycle, it is feasible that trimerization is an important strategy to capture special lipid components that are relevant to the protein activity.
- Department of Physics, Graduate School of Science, Nagoya University, Chikusa-ku, Nagoya 464-8602, Japan.
Organizational Affiliation: