Structure-Based Design, Synthesis, and Evaluation of Peptide-Mimetic SARS 3CL Protease Inhibitors.
Akaji, K., Konno, H., Mitsui, H., Teruya, K., Shimamoto, Y., Hattori, Y., Ozaki, T., Kusunoki, M., Sanjoh, A.(2011) J Med Chem 54: 7962-7973
- PubMed: 22014094 Search on PubMed
- DOI: https://doi.org/10.1021/jm200870n
- Primary Citation Related Structures:
3ATW, 3AVZ, 3AW0, 3AW1 - PubMed Abstract:
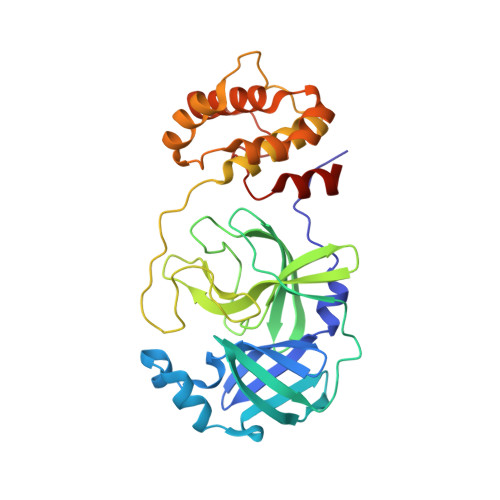
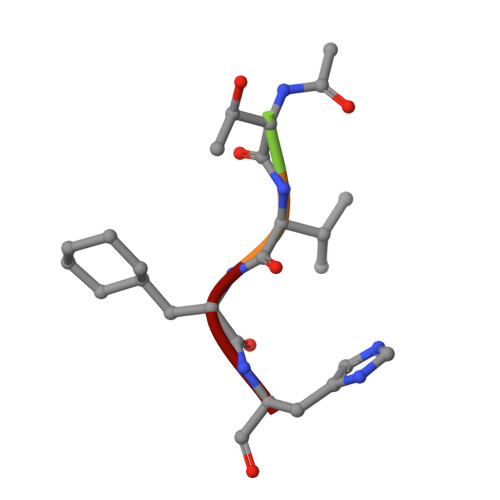
The design and evaluation of low molecular weight peptide-based severe acute respiratory syndrome (SARS) chymotrypsin-like protease (3CL) protease inhibitors are described. A substrate-based peptide aldehyde was selected as a starting compound, and optimum side-chain structures were determined, based on a comparison of inhibitory activities with Michael type inhibitors. For the efficient screening of peptide aldehydes containing a specific C-terminal residue, a new approach employing thioacetal to aldehyde conversion mediated by N-bromosuccinimide was devised. Structural optimization was carried out based on X-ray crystallographic analyses of the R188I SARS 3CL protease in a complex with each inhibitor to provide a tetrapeptide aldehyde with an IC(50) value of 98 nM. The resulting compound carried no substrate sequence, except for a P(3) site directed toward the outside of the protease. X-ray crystallography provided insights into the protein-ligand interactions.
- Department of Medicinal Chemistry, Kyoto Pharmaceutical University, Yamashina-ku, Kyoto 607-8412, Japan.
Organizational Affiliation: