The Development of N-alpha-(2-Carboxyl)benzoyl-N(5)-(2-fluoro-1-iminoethyl)-l-ornithine Amide (o-F-amidine) and N-alpha-(2-Carboxyl)benzoyl-N(5)-(2-chloro-1-iminoethyl)-l-ornithine Amide (o-Cl-amidine) As Second Generation Protein Arginine Deiminase (PAD) Inhibitors
Causey, C.P., Jones, J.E., Slack, J.L., Kamei, D., Jones Jr, L.E., Subramanian, V., Knuckley, B., Ebrahimi, P., Chumanevich, A.A., Luo, Y., Hashimoto, H., Sato, M., Hofseth, L.J., Thompson, P.R.(2011) J Med Chem 54: 6919-6935
- PubMed: 21882827
- DOI: https://doi.org/10.1021/jm2008985
- Primary Citation of Related Structures:
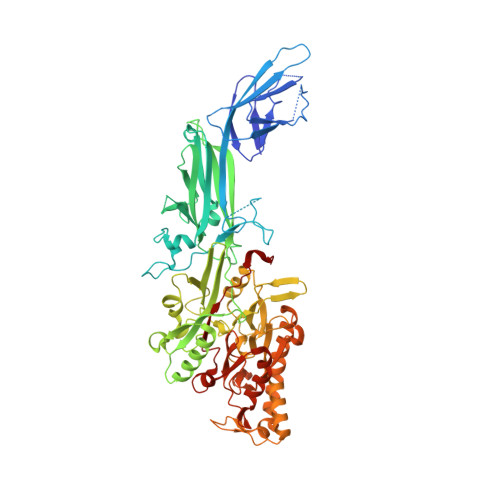
3B1T, 3B1U - PubMed Abstract:
Protein arginine deiminase (PAD) activity is upregulated in a number of human diseases, including rheumatoid arthritis, ulcerative colitis, and cancer. These enzymes, there are five in humans (PADs 1-4 and 6), regulate gene transcription, cellular differentiation, and the innate immune response. Building on our successful generation of F- and Cl-amidine, which irreversibly inhibit all of the PADs, a structure-activity relationship was performed to develop second generation compounds with improved potency and selectivity. Incorporation of a carboxylate ortho to the backbone amide resulted in the identification of N-α-(2-carboxyl)benzoyl-N(5)-(2-fluoro-1-iminoethyl)-l-ornithine amide (o-F-amidine) and N-α-(2-carboxyl)benzoyl-N(5)-(2-chloro-1-iminoethyl)-l-ornithine amide (o-Cl-amidine), as PAD inactivators with improved potency (up to 65-fold) and selectivity (up to 25-fold). Relative to F- and Cl-amidine, the compounds also show enhanced potency in cellulo. As such, these compounds will be versatile chemical probes of PAD function.
- Department of Chemistry, The Scripps Research Institute, 130 Scripps Way, Jupiter, Florida 33458, United States.
Organizational Affiliation: