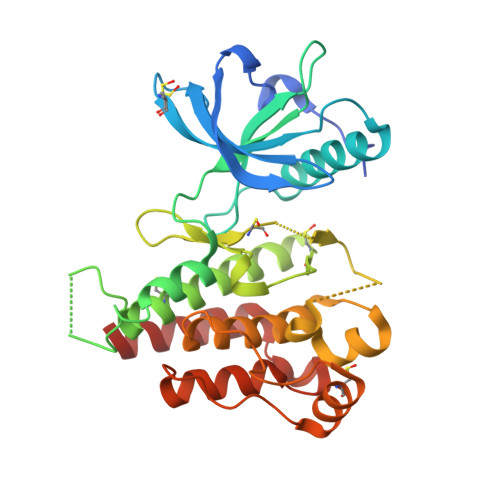
Discovery of 5-[[4-[(2,3-dimethyl-2H-indazol-6-yl)methylamino]-2-pyrimidinyl]amino]-2-methyl-benzenesulfonamide (Pazopanib), a novel and potent vascular endothelial growth factor receptor inhibitor.
Harris, P.A., Boloor, A., Cheung, M., Kumar, R., Crosby, R.M., Davis-Ward, R.G., Epperly, A.H., Hinkle, K.W., Hunter, R.N., Johnson, J.H., Knick, V.B., Laudeman, C.P., Luttrell, D.K., Mook, R.A., Nolte, R.T., Rudolph, S.K., Szewczyk, J.R., Truesdale, A.T., Veal, J.M., Wang, L., Stafford, J.A.(2008) J Med Chem 51: 4632-4640
- PubMed: 18620382 Search on PubMed
- DOI: https://doi.org/10.1021/jm800566m
- Primary Citation Related Structures:
3CJF, 3CJG - PubMed Abstract:
Inhibition of the vascular endothelial growth factor (VEGF) signaling pathway has emerged as one of the most promising new approaches for cancer therapy. We describe herein the key steps starting from an initial screening hit leading to the discovery of pazopanib, N(4)-(2,3-dimethyl-2H-indazol-6-yl)-N(4)-methyl-N(2)-(4-methyl-3-sulfonamidophenyl)-2,4-pyrimidinediamine, a potent pan-VEGF receptor (VEGFR) inhibitor under clinical development for renal-cell cancer and other solid tumors.
- GlaxoSmithKline, Five Moore Drive, Research Triangle Park, North Carolina 27709, USA. philip.a.harris@gsk.com
Organizational Affiliation: