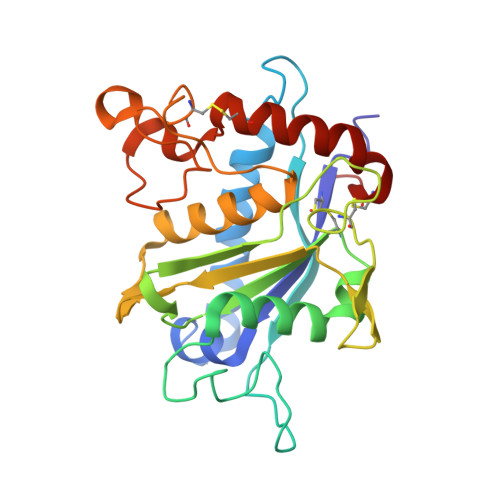
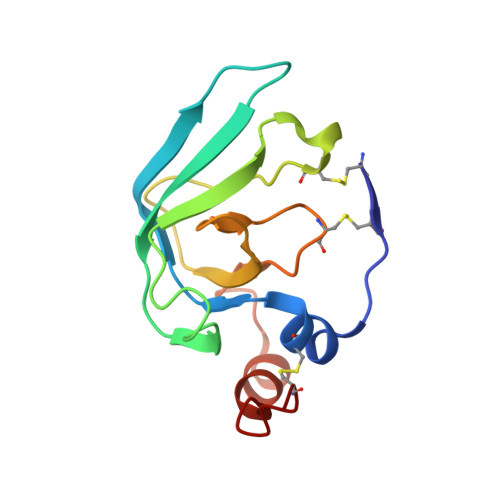
Structural determinants of the ADAM inhibition by TIMP-3: crystal structure of the TACE-N-TIMP-3 complex.
Wisniewska, M., Goettig, P., Maskos, K., Belouski, E., Winters, D., Hecht, R., Black, R., Bode, W.(2008) J Mol Biology 381: 1307-1319
- PubMed: 18638486 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.06.088
- Primary Citation Related Structures:
3CKI - PubMed Abstract:
TIMP-3 (tissue inhibitor of metalloproteinases 3) is unique among the TIMP inhibitors, in that it effectively inhibits the TNF-alpha converting enzyme (TACE). In order to understand this selective capability of inhibition, we crystallized the complex formed by the catalytic domain of recombinant human TACE and the N-terminal domain of TIMP-3 (N-TIMP-3), and determined its molecular structure with X-ray data to 2.3 A resolution. The structure reveals that TIMP-3 exhibits a fold similar to those of TIMP-1 and TIMP-2, and interacts through its functional binding edge, which consists of the N-terminal segment and other loops, with the active-site cleft of TACE in a manner similar to that of matrix metalloproteinases (MMPs). Therefore, the mechanism of TIMP-3 binding toward TACE is not fundamentally different from that previously elucidated for the MMPs. The Phe34 phenyl side chain situated at the tip of the relatively short sA-sB loop of TIMP-3 extends into a unique hydrophobic groove of the TACE surface, and two Leu residues in the adjacent sC-connector and sE-sF loops are tightly packed in the interface allowing favourable interactions, in agreement with predictions obtained by systematic mutations by Gillian Murphy's group. The combination of favourable functional epitopes together with a considerable flexibility renders TIMP-3 an efficient TACE inhibitor. This structure might provide means to design more efficient TIMP inhibitors of TACE.
- Max-Planck-Institut für Biochemie, Proteinase Research Group, Am Klopferspitz 18, D-82152 Martinsried, Germany.
Organizational Affiliation: