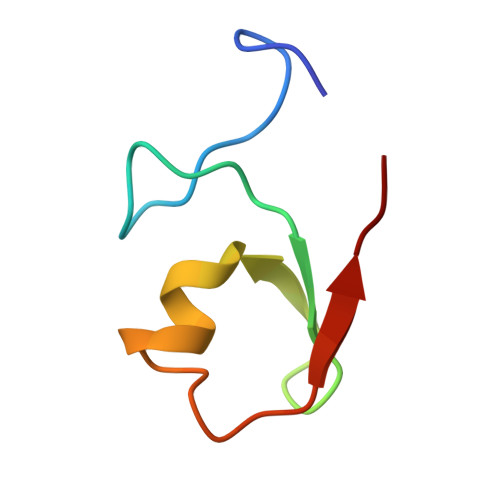
Structural analysis of B-Box 2 from MuRF1: identification of a novel self-association pattern in a RING-like fold
Mrosek, M., Meier, S., Ucurum-Fotiadis, Z., von Castelmur, E., Hedbom, E., Lustig, A., Grzesiek, S., Labeit, D., Labeit, S., Mayans, O.(2008) Biochemistry 47: 10722-10730
- PubMed: 18795805 Search on PubMed
- DOI: https://doi.org/10.1021/bi800733z
- Primary Citation Related Structures:
3DDT - PubMed Abstract:
The B-box motif is the defining feature of the TRIM family of proteins, characterized by a RING finger-B-box-coiled coil tripartite fold. We have elucidated the crystal structure of B-box 2 (B2) from MuRF1, a TRIM protein that supports a wide variety of protein interactions in the sarcomere and regulates the trophic state of striated muscle tissue. MuRF1 B2 coordinates two zinc ions through a cross-brace alpha/beta-topology typical of members of the RING finger superfamily. However, it self-associates into dimers with high affinity. The dimerization pattern is mediated by the helical component of this fold and is unique among RING-like folds. This B2 reveals a long shallow groove that encircles the C-terminal metal binding site ZnII and appears as the defining protein-protein interaction feature of this domain. A cluster of conserved hydrophobic residues in this groove and, in particular, a highly conserved aromatic residue (Y133 in MuRF1 B2) is likely to be central to this role. We expect these findings to aid the future exploration of the cellular function and therapeutic potential of MuRF1.
- Division of Structural Biology, Biozentrum, University of Basel, Klingelbergstrasse 70, CH-4056 Basel, Switzerland.
Organizational Affiliation: