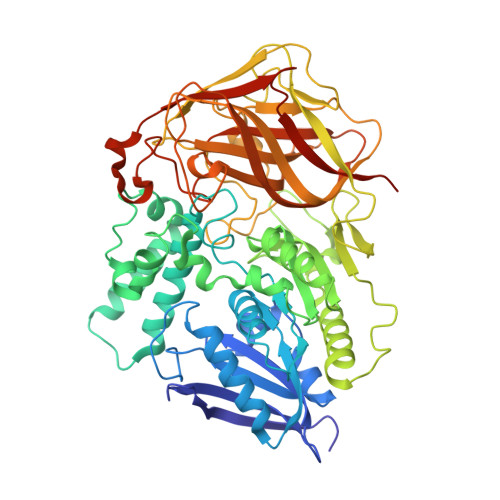
Structural analysis of thermostabilizing mutations of cocaine esterase.
Narasimhan, D., Nance, M.R., Gao, D., Ko, M.C., Macdonald, J., Tamburi, P., Yoon, D., Landry, D.M., Woods, J.H., Zhan, C.G., Tesmer, J.J., Sunahara, R.K.(2010) Protein Eng Des Sel 23: 537-547
- PubMed: 20436035 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/protein/gzq025
- Primary Citation Related Structures:
3I2F, 3I2G, 3I2H, 3I2I, 3I2J, 3I2K - PubMed Abstract:
Cocaine is considered to be the most addictive of all substances of abuse and mediates its effects by inhibiting monoamine transporters, primarily the dopamine transporters. There are currently no small molecules that can be used to combat its toxic and addictive properties, in part because of the difficulty of developing compounds that inhibit cocaine binding without having intrinsic effects on dopamine transport. Most of the effective cocaine inhibitors also display addictive properties. We have recently reported the use of cocaine esterase (CocE) to accelerate the removal of systemic cocaine and to prevent cocaine-induced lethality. However, wild-type CocE is relatively unstable at physiological temperatures (tau(1/2) approximately 13 min at 37 degrees C), presenting challenges for its development as a viable therapeutic agent. We applied computational approaches to predict mutations to stabilize CocE and showed that several of these have increased stability both in vitro and in vivo, with the most efficacious mutant (T172R/G173Q) extending half-life up to 370 min. Here we present novel X-ray crystallographic data on these mutants that provide a plausible model for the observed enhanced stability. We also more extensively characterize the previously reported variants and report on a new stabilizing mutant, L169K. The improved stability of these engineered CocE enzymes will have a profound influence on the use of this protein to combat cocaine-induced toxicity and addiction in humans.
- Department of Pharmacology, University of Michigan, Ann Arbor, MI, USA.
Organizational Affiliation: