Synthesis and biological activities of novel indole derivatives as potent and selective PPAR-gamma modulators
Lamotte, Y., Martres, P., Faucher, N., Laroze, A., Grillot, D., Ancellin, N., Saintillan, Y., Beneton, V., Gampe, R.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Peroxisome proliferator-activated receptor gamma | A, C [auth D] | 272 | Homo sapiens | Mutation(s): 0 Gene Names: PPARG, NR1C3 | 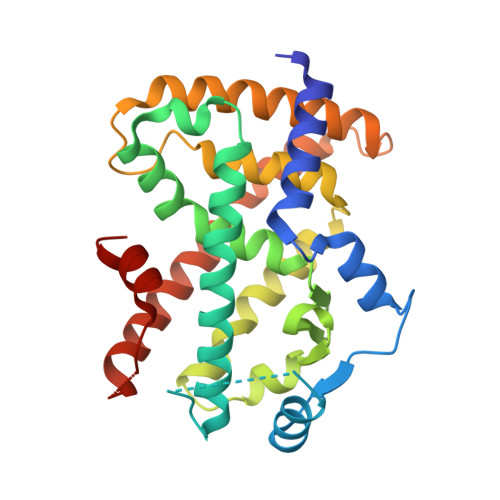 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P37231 GTEx: ENSG00000132170 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P37231 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Steroid Receptor Coactivator-1 | B, D [auth E] | 27 | N/A | Mutation(s): 0 | 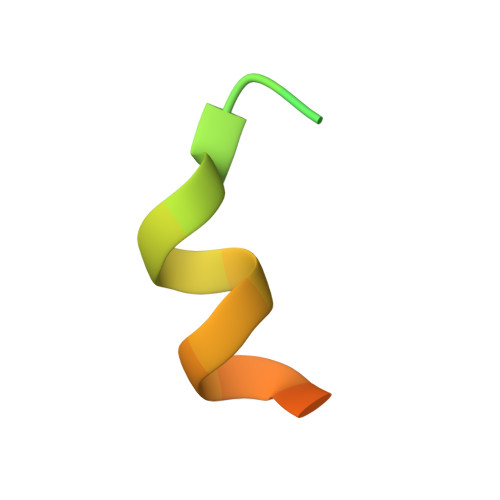 |
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 538 Download:Ideal Coordinates CCD File | E [auth A], F [auth D] | 4'-[(2,3-dimethyl-5-{[(1S)-1-phenylpropyl]carbamoyl}-1H-indol-1-yl)methyl]biphenyl-2-carboxylic acid C34 H32 N2 O3 GAGNYMUXGIUCTR-HKBQPEDESA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 83.335 | α = 90 |
| b = 84.158 | β = 90 |
| c = 96.425 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| JDirector | data collection |
| MOLREP | phasing |