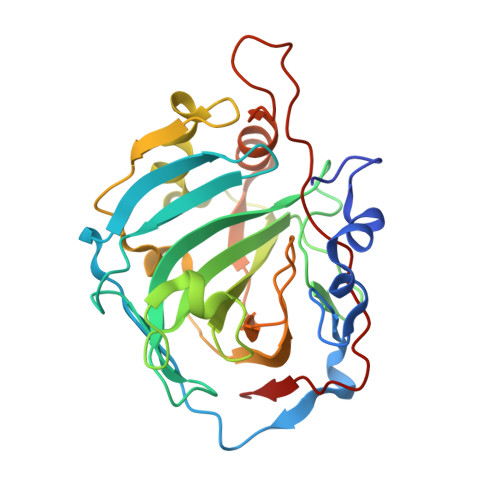
The first example of a significant active site conformational rearrangement in a carbonic anhydrase-inhibitor adduct: the carbonic anhydrase I-topiramate complex.
Alterio, V., Monti, S.M., Truppo, E., Pedone, C., Supuran, C.T., De Simone, G.(2010) Org Biomol Chem
- PubMed: 20505865 Search on PubMed
- DOI: https://doi.org/10.1039/b926832d
- Primary Citation Related Structures:
3LXE - PubMed Abstract:
Topiramate is a widely used antiepileptic drug, which has been demonstrated to act as an efficient weight loss agent. Since several studies have pointed out that is a potent in vitro inhibitor of several Carbonic anhydrase (CA) isozymes, it has been hypothesized that its anti-obesity properties could be ascribed to the inhibition of the CAs involved in de novo lipogenesis. Consequently, the study of the interactions of with all human CA isoforms represents an important step for the rational drug design of selective CA inhibitors to be used as anti-obesity drugs. In this paper we report the crystallographic structure of the adduct that forms with hCA I, showing for the first time a profound reorganization of the CA active site upon binding of the inhibitor. Moreover, a structural comparison with hCA II- and hCA VA- adducts, previously investigated, has been performed showing that a different H-bond network together with the movement of some amino acid residues in the active site may account for the different inhibition constants of toward these three CA isozymes.
- Istituto di Biostrutture e Bioimmagini-CNR, via Mezzocannone 16, 80134 Naples, Italy.
Organizational Affiliation: