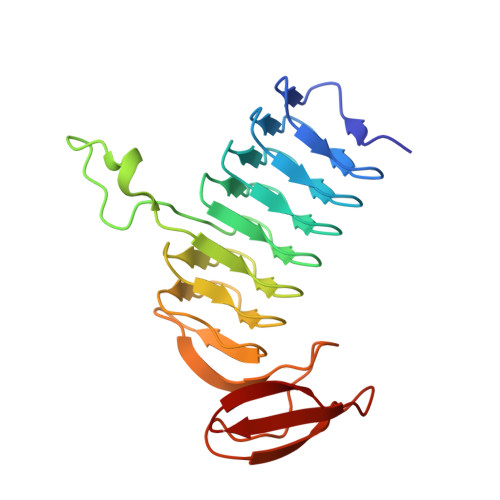
Molecular structure of WlbB, a bacterial N-acetyltransferase involved in the biosynthesis of 2,3-diacetamido-2,3-dideoxy-D-mannuronic acid .
Thoden, J.B., Holden, H.M.(2010) Biochemistry 49: 4644-4653
- PubMed: 20433200
- DOI: https://doi.org/10.1021/bi1005738
- Primary Citation Related Structures:
3MQG, 3MQH - PubMed Abstract:
The pathogenic bacteria Pseudomonas aeruginosa and Bordetella pertussis contain in their outer membranes the rare sugar 2,3-diacetamido-2,3-dideoxy-d-mannuronic acid. Five enzymes are required for the biosynthesis of this sugar starting from UDP-N-acetylglucosamine. One of these, referred to as WlbB, is an N-acetyltransferase that converts UDP-2-acetamido-3-amino-2,3-dideoxy-d-glucuronic acid (UDP-GlcNAc3NA) to UDP-2,3-diacetamido-2,3-dideoxy-d-glucuronic acid (UDP-GlcNAc3NAcA). Here we report the three-dimensional structure of WlbB from Bordetella petrii. For this analysis, two ternary structures were determined to 1.43 A resolution: one in which the protein was complexed with acetyl-CoA and UDP and the second in which the protein contained bound CoA and UDP-GlcNAc3NA. WlbB adopts a trimeric quaternary structure and belongs to the LbetaH superfamily of N-acyltransferases. Each subunit contains 27 beta-strands, 23 of which form the canonical left-handed beta-helix. There are only two hydrogen bonds that occur between the protein and the GlcNAc3NA moiety, one between O(delta1) of Asn 84 and the sugar C-3' amino group and the second between the backbone amide group of Arg 94 and the sugar C-5' carboxylate. The sugar C-3' amino group is ideally positioned in the active site to attack the si face of acetyl-CoA. Given that there are no protein side chains that can function as general bases within the GlcNAc3NA binding pocket, a reaction mechanism is proposed for WlbB whereby the sulfur of CoA ultimately functions as the proton acceptor required for catalysis.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: