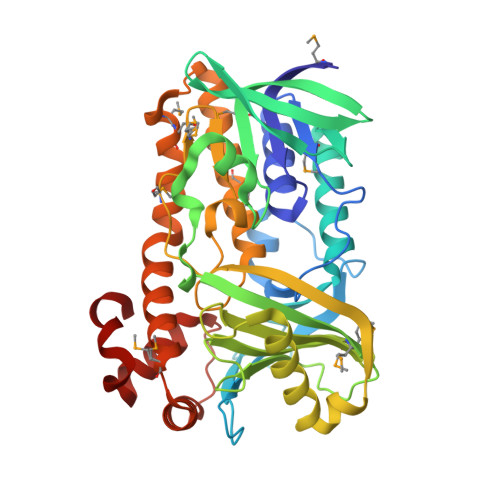
Insights into substrate specificity of geranylgeranyl reductases revealed by the structure of digeranylgeranylglycerophospholipid reductase, an essential enzyme in the biosynthesis of archaeal membrane lipids.
Xu, Q., Eguchi, T., Mathews, I.I., Rife, C.L., Chiu, H.J., Farr, C.L., Feuerhelm, J., Jaroszewski, L., Klock, H.E., Knuth, M.W., Miller, M.D., Weekes, D., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) J Mol Biology 404: 403-417
- PubMed: 20869368
- DOI: https://doi.org/10.1016/j.jmb.2010.09.032
- Primary Citation of Related Structures:
3OZ2 - PubMed Abstract:
Archaeal membrane lipids consist of branched, saturated hydrocarbons distinct from those found in bacteria and eukaryotes. Digeranylgeranylglycerophospholipid reductase (DGGR) catalyzes the hydrogenation process that converts unsaturated 2,3-di-O-geranylgeranylglyceryl phosphate to saturated 2,3-di-O-phytanylglyceryl phosphate as a critical step in the biosynthesis of archaeal membrane lipids. The saturation of hydrocarbon chains confers the ability to resist hydrolysis and oxidation and helps archaea withstand extreme conditions. DGGR is a member of the geranylgeranyl reductase family that is also widely distributed in bacteria and plants, where the family members are involved in the biosynthesis of photosynthetic pigments. We have determined the crystal structure of DGGR from the thermophilic heterotrophic archaea Thermoplasma acidophilum at 1.6 Å resolution, in complex with flavin adenine dinucleotide (FAD) and a bacterial lipid. The DGGR structure can be assigned to the well-studied, p-hydroxybenzoate hydroxylase (PHBH) SCOP superfamily of flavoproteins that include many aromatic hydroxylases and other enzymes with diverse functions. In the DGGR complex, FAD adopts the IN conformation (closed) previously observed in other PHBH flavoproteins. DGGR contains a large substrate-binding site that extends across the entire ligand-binding domain. Electron density corresponding to a bacterial lipid was found within this cavity. The cavity consists of a large opening that tapers down to two, narrow, curved tunnels that closely mimic the shape of the preferred substrate. We identified a sequence motif, PxxYxWxFP, that defines a specificity pocket in the enzyme and precisely aligns the double bond of the geranyl group with respect to the FAD cofactor, thus providing a structural basis for the substrate specificity of geranylgeranyl reductases. DGGR is likely to share a common mechanism with other PHBH enzymes in which FAD switches between two conformations that correspond to the reductive and oxidative half cycles. The structure provides evidence that substrate binding likely involves conformational changes, which are coupled to the two conformational states of the FAD.
- Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA 94025, USA.
Organizational Affiliation: