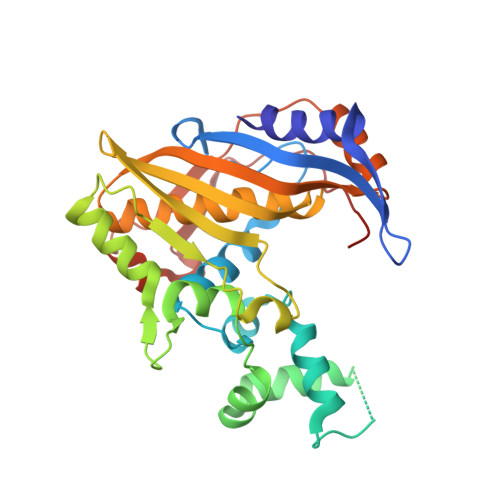
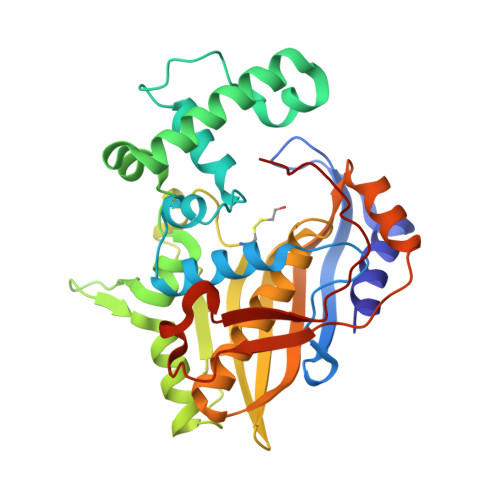
The structure of Enterococcus faecalis thymidylate synthase provides clues about folate bacterial metabolism.
Pozzi, C., Ferrari, S., Cortesi, D., Luciani, R., Stroud, R.M., Catalano, A., Costi, M.P., Mangani, S.(2012) Acta Crystallogr D Biol Crystallogr 68: 1232-1241
- PubMed: 22948925 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444912026236
- Primary Citation Related Structures:
3UWL - PubMed Abstract:
Drug resistance to therapeutic antibiotics poses a challenge to the identification of novel targets and drugs for the treatment of infectious diseases. Infections caused by Enterococcus faecalis are a major health problem. Thymidylate synthase (TS) from E. faecalis is a potential target for antibacterial therapy. The X-ray crystallographic structure of E. faecalis thymidylate synthase (EfTS), which was obtained as a native binary complex composed of EfTS and 5-formyltetrahydrofolate (5-FTHF), has been determined. The structure provides evidence that EfTS is a half-of-the-sites reactive enzyme, as 5-FTHF is bound to two of the four independent subunits present in the crystal asymmetric unit. 5-FTHF is a metabolite of the one-carbon transfer reaction catalysed by 5-formyltetrahydrofolate cyclo-ligase. Kinetic studies show that 5-FTHF is a weak inhibitor of EfTS, suggesting that the EfTS-5-FTHF complex may function as a source of folates and/or may regulate one-carbon metabolism. The structure represents the first example of endogenous 5-FTHF bound to a protein involved in folate metabolism.
- Dipartimento di Chimica, University of Siena, Siena, Italy.
Organizational Affiliation: